Data Visualization Using R
Deepayan Sarkar
R graphics
R comes with two largely independent graphics subsystems
“Traditional” graphics (package
graphics)- Available in R from the beginning
- Rich collection of tools
- Not very flexible
Grid graphics (package
grid)- Relatively recent (2000)
- Low-level tool, highly flexible
Grid forms the basis of two high-level graphics systems:
- Package
lattice: based on Trellis graphics (Cleveland) - Package
ggplot2: inspired by “Grammar of Graphics” (Wilkinson)
- Package
R graphics
It is also possible to interface with external graphics systems.
This is useful when some kind of interaction is required.
Package
rgl: Interactive 3-D plots with OpenGLPackage
plotly: Javascript-based plots in browserPackage
rggobi: Interactive and dynamic graphics using GGobi
We will see a little bit of all these.
Traditional graphics - some history
Like the language itself, R graphics was derived from S (Bell Labs, 1970s)
S graphics was based on the GRZ model:
May be described as a “painter’s model”
Graphic is built out of “primitives” such as line segments, polygons, text, etc.
Later elements are drawn on top of earlier ones
No provision for deleting an element once it was drawn
This allows graphics output to be easily abstracted
Output devices: screen, PDF, PNG
Enough to implement primitives for each device
Also impacted how plots were constructed
Consequence of the painter’s model
Mental approach: a plot is a work-in-progress
Always possibile to add something more
This attitude pervades traditional graphics
- Typical approach:
- Start with a more or less complete (high-level) plot
- Add (low-level) elements to customize
An example: Anscombe’s dataset 1
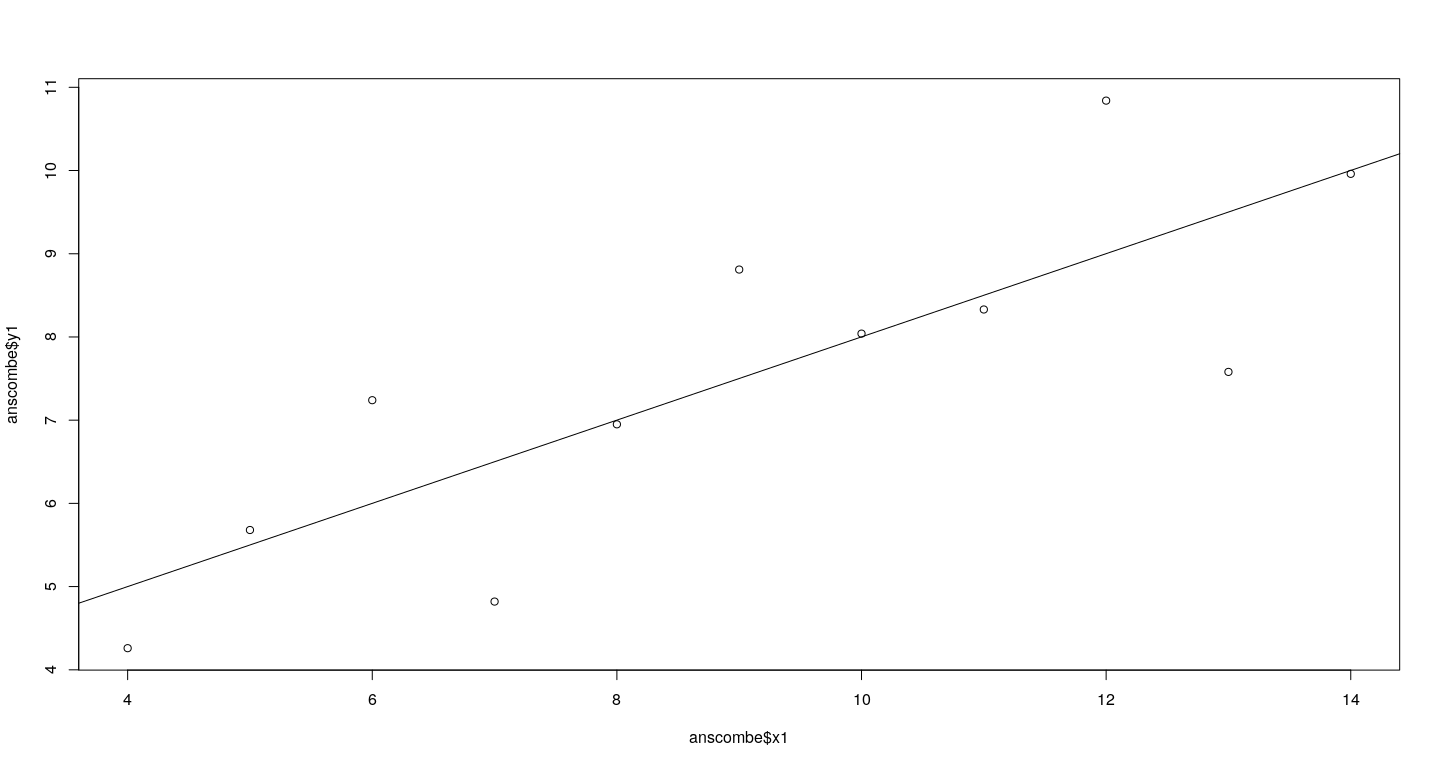
An example: Anscombe’s dataset 2
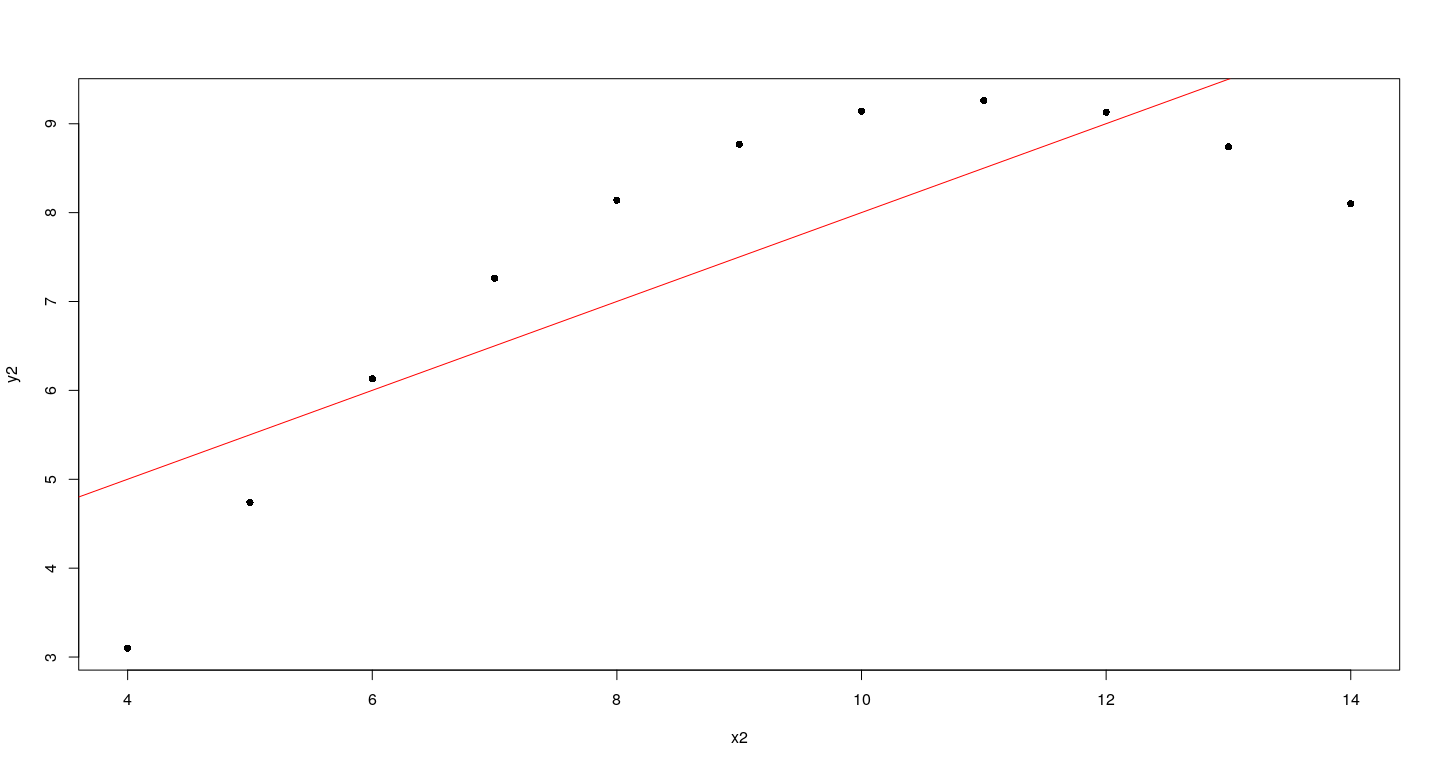
How could this plot have been created using low-level functions?
Try running this code one line at a time:
plot(anscombe$x1, anscombe$y1, type = "n", axes = FALSE, xlab = "", ylab = "")
box()
points(anscombe$x1, anscombe$y1, pch = 16)
axis(side = 1)
axis(side = 2)
title(main = "Anscombe's first dataset", xlab = "x1", ylab = "y1")
abline(lm(y1 ~ x1, anscombe), col = "red")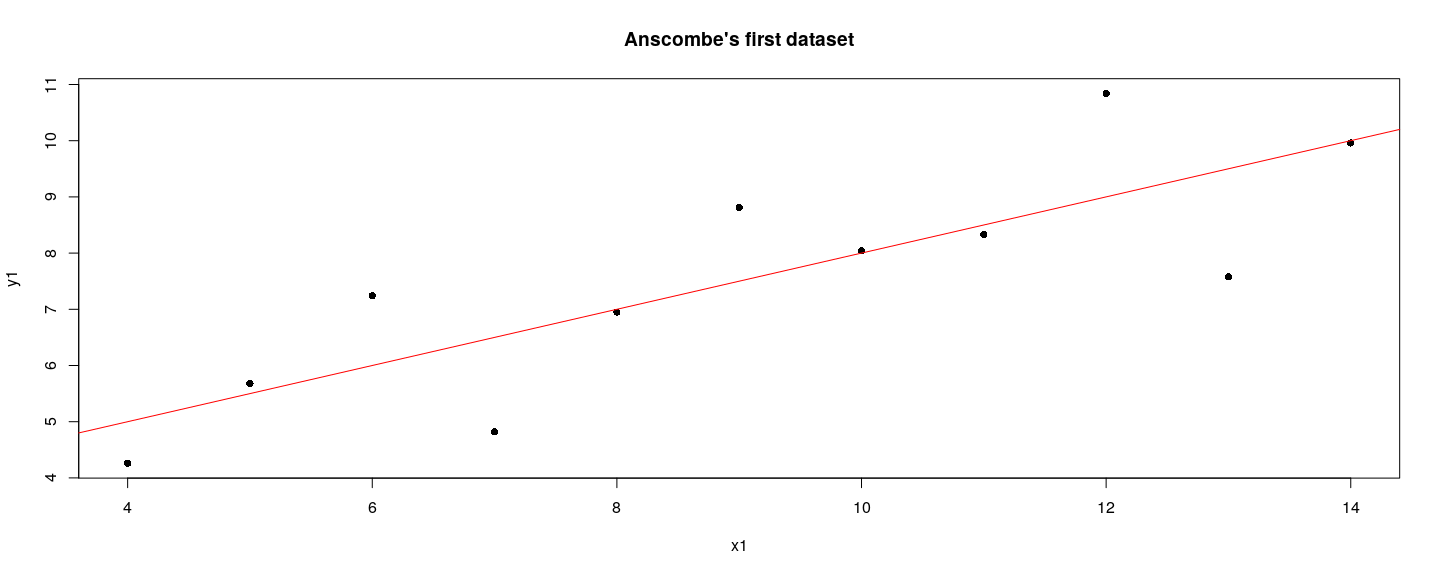
A more polished version
plot(anscombe$x2, anscombe$y2, type = "n", axes = FALSE, xlab = "", ylab = "")
lims <- par("usr")
rect(lims[1], lims[3], lims[2], lims[4], col = "grey80", border = NA)
abline(v = pretty(lims[1:2]), h = pretty(lims[3:4]), col = "white", lwd = 2)
axis(side = 1, col = "grey80", col.axis = "grey20")
axis(side = 2, col = "grey80", col.axis = "grey20", las = 1)
points(anscombe$x2, anscombe$y2, pch = 16)
title(main = "Anscombe's second dataset", xlab = "x2", ylab = "y2", col = "grey20")
abline(lm(y2 ~ x2, anscombe), col = "grey20")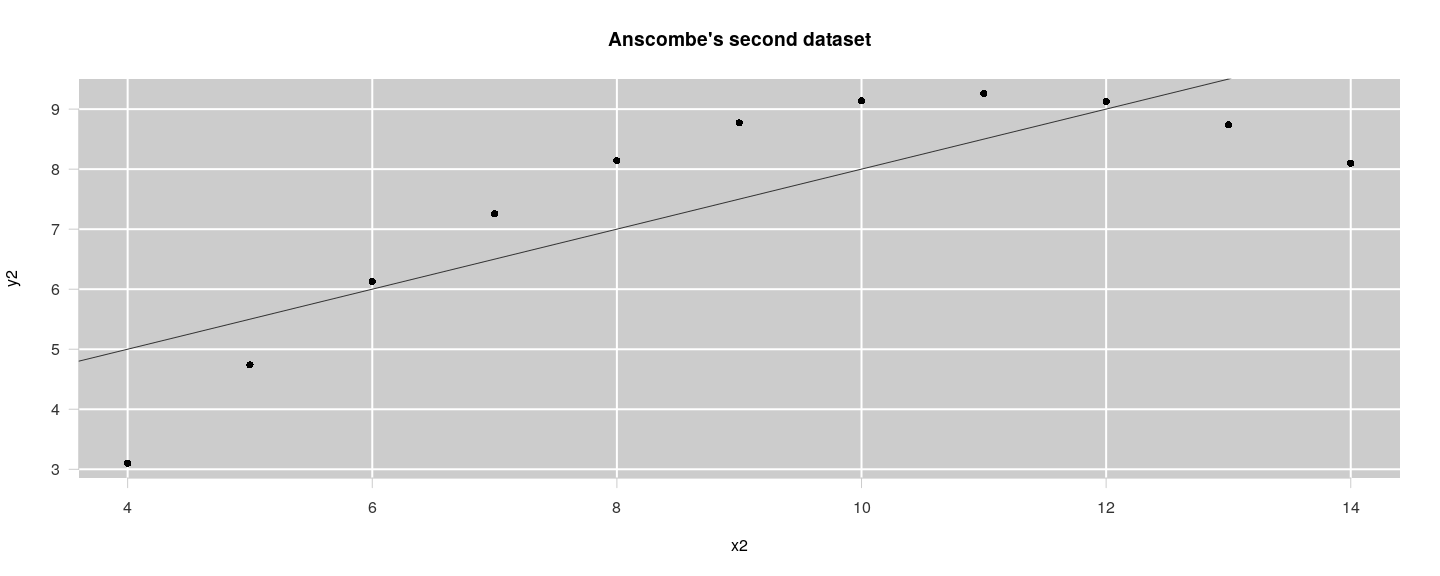
Other high-level plots
Generally, traditional graphics work by calling specialized high-level functions
- Some you have already learned about:
- Histograms / density plots
- Q-Q plots
- Dot plots
- Bar charts
- Box and whisker plots
Let’s see some more examples using the
airqualitydataset.
'data.frame': 153 obs. of 6 variables:
$ Ozone : int 41 36 12 18 NA 28 23 19 8 NA ...
$ Solar.R: int 190 118 149 313 NA NA 299 99 19 194 ...
$ Wind : num 7.4 8 12.6 11.5 14.3 14.9 8.6 13.8 20.1 8.6 ...
$ Temp : int 67 72 74 62 56 66 65 59 61 69 ...
$ Month : int 5 5 5 5 5 5 5 5 5 5 ...
$ Day : int 1 2 3 4 5 6 7 8 9 10 ...Other high-level plots - examples
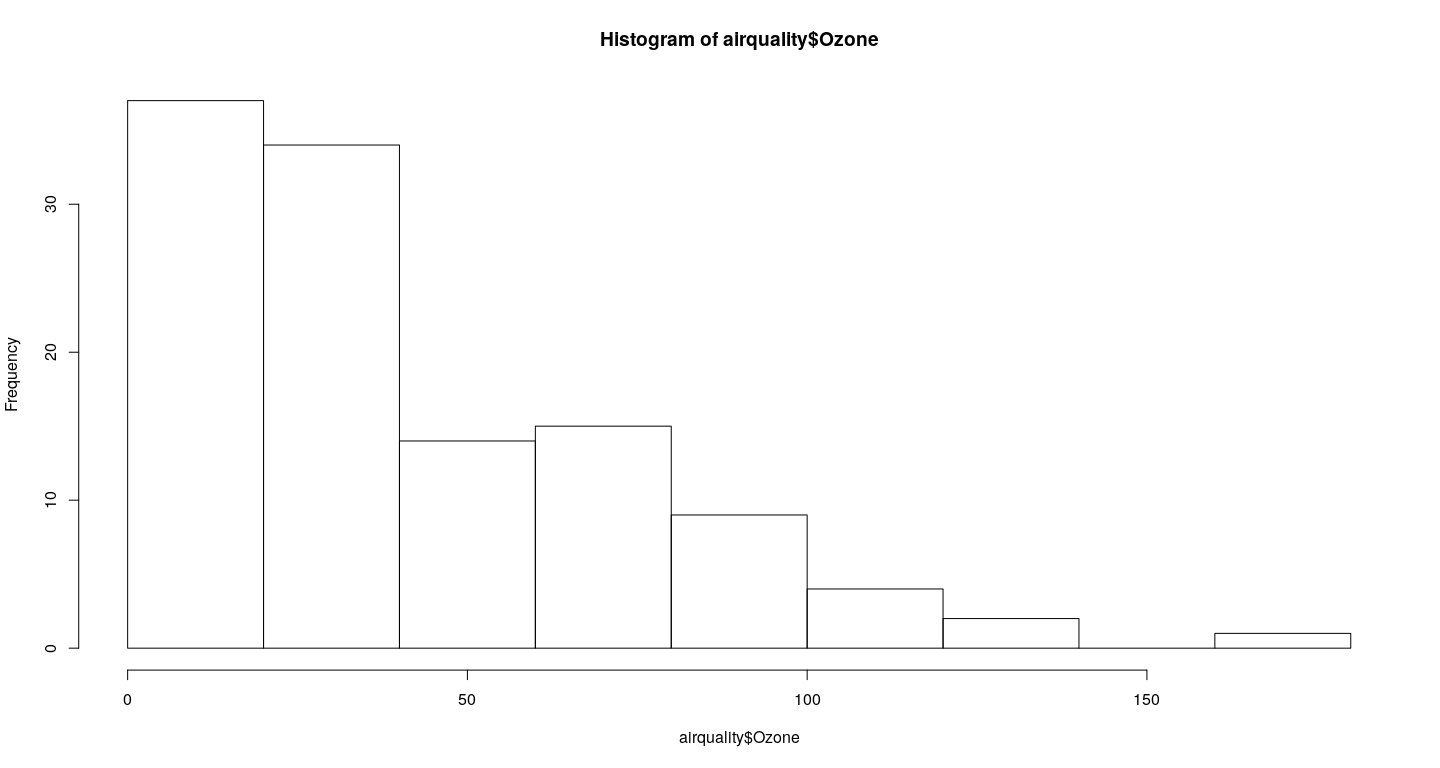
Other high-level plots - examples
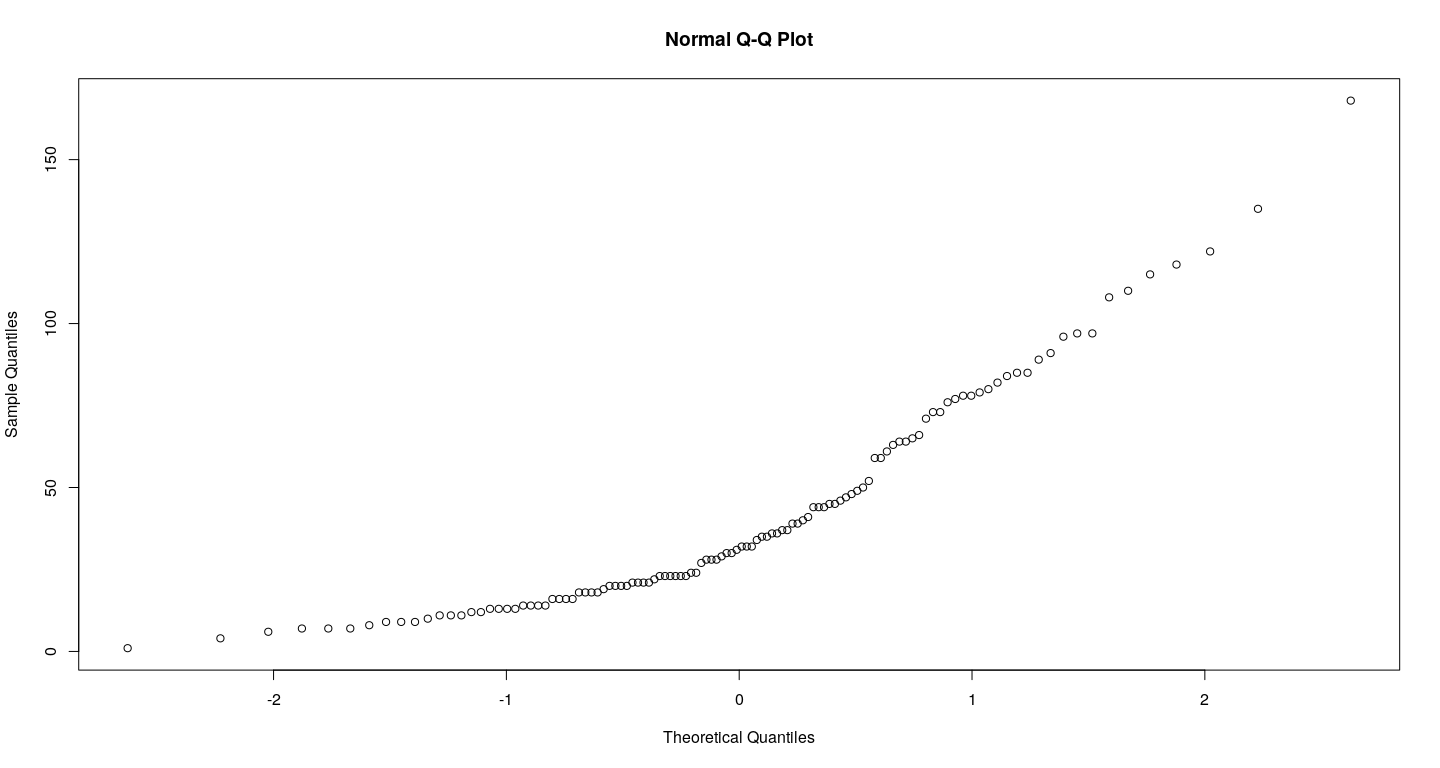
Other high-level plots - examples
airquality$fmonth <- with(airquality, factor(Month, levels = 1:12, labels = month.name))
boxplot(Ozone ~ droplevels(fmonth), data = airquality)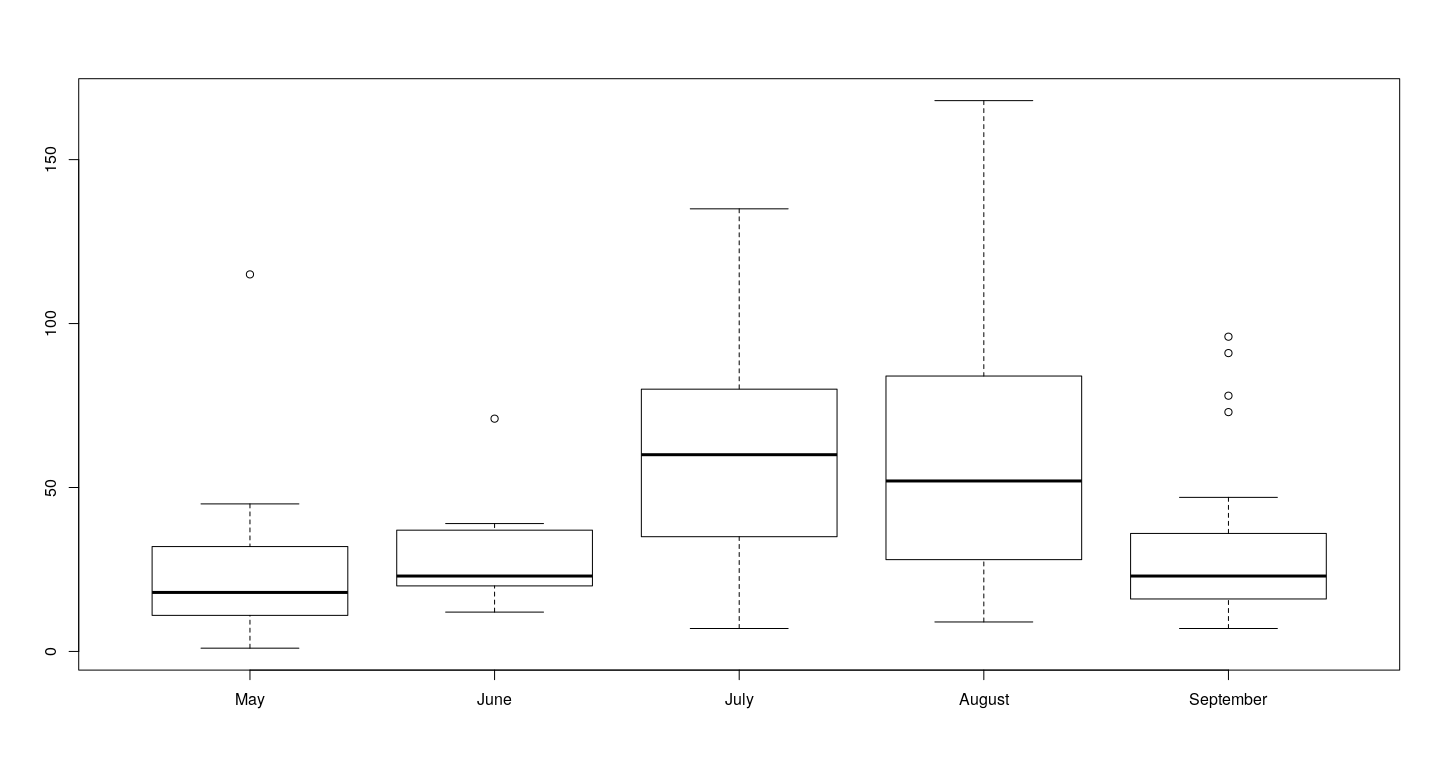
Comparing subsets
The last example (box and whisker plot) is different — it allows comparison!
Making comparisons is one of the primary goals of statistical graphics
How can we compare other plots like scatter plots and histograms?
- Two solutions: juxtaposition or superposition
Juxtaposing by splitting figure region
par(mfrow = c(2, 2))
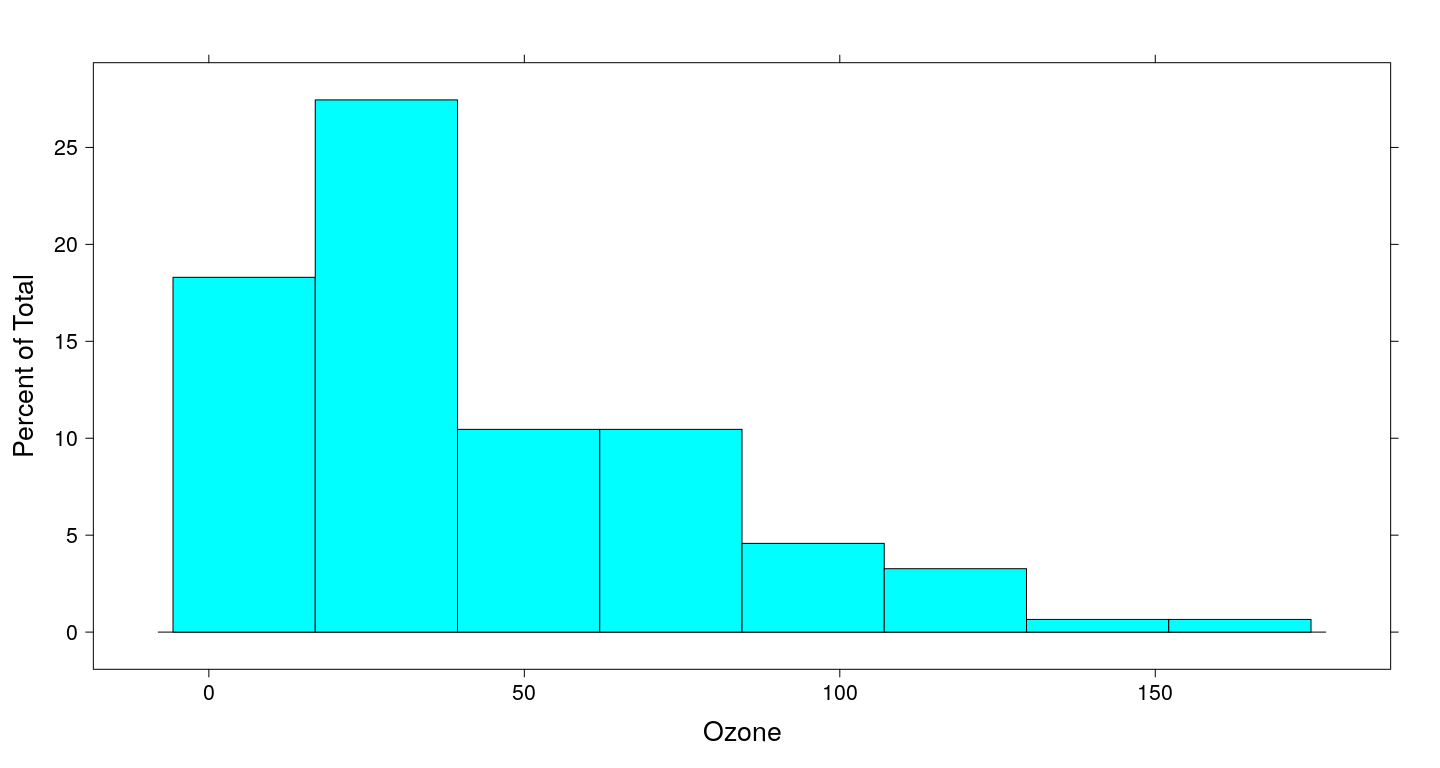
hist(airquality$Ozone)
hist(airquality$Solar.R)
hist(airquality$Wind)
hist(airquality$Temp)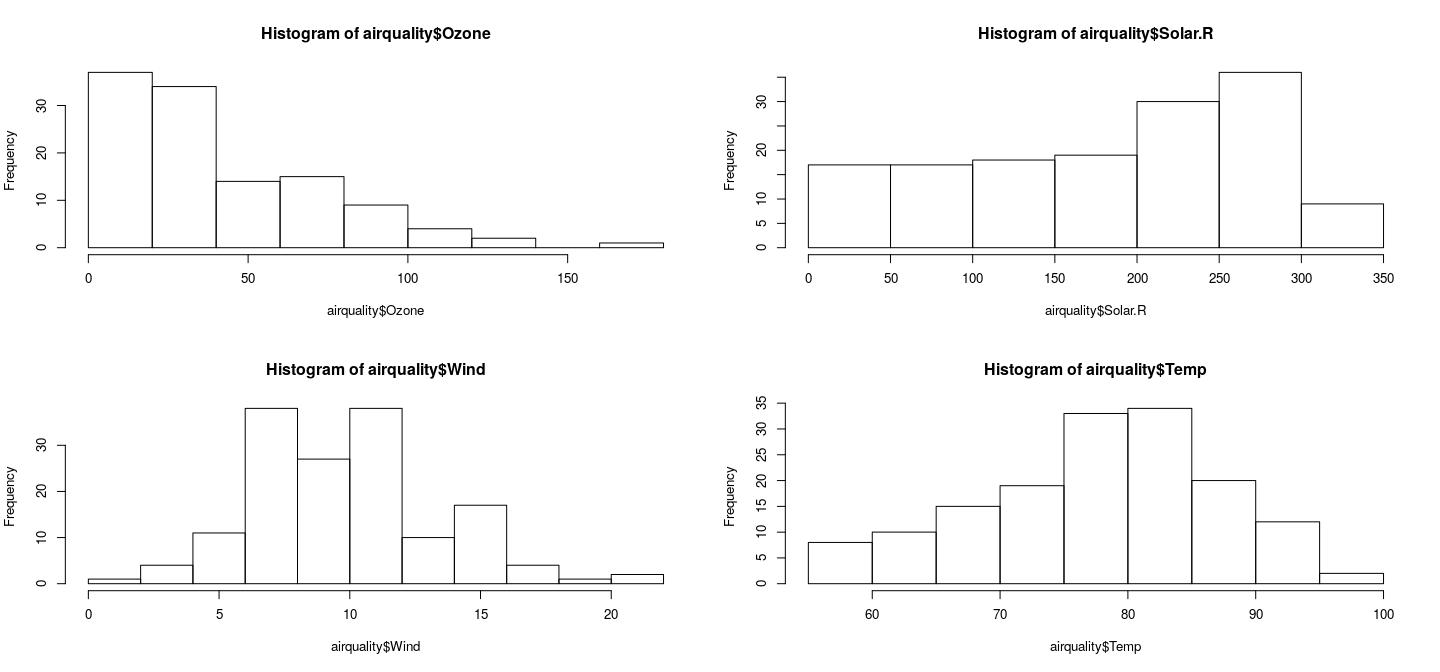
Juxtaposing by splitting figure region
List of 5
$ May : int [1:31] 67 72 74 62 56 66 65 59 61 69 ...
$ June : int [1:30] 78 74 67 84 85 79 82 87 90 87 ...
$ July : int [1:31] 84 85 81 84 83 83 88 92 92 89 ...
$ August : int [1:31] 81 81 82 86 85 87 89 90 90 92 ...
$ September: int [1:30] 91 92 93 93 87 84 80 78 75 73 ...Juxtaposing by splitting figure region
par(mfrow = c(2, 3))
qqnorm(s$May, main = "May")
qqnorm(s$June, main = "June")
qqnorm(s$July, main = "July")
qqnorm(s$August, main = "August")
qqnorm(s$September, main = "September")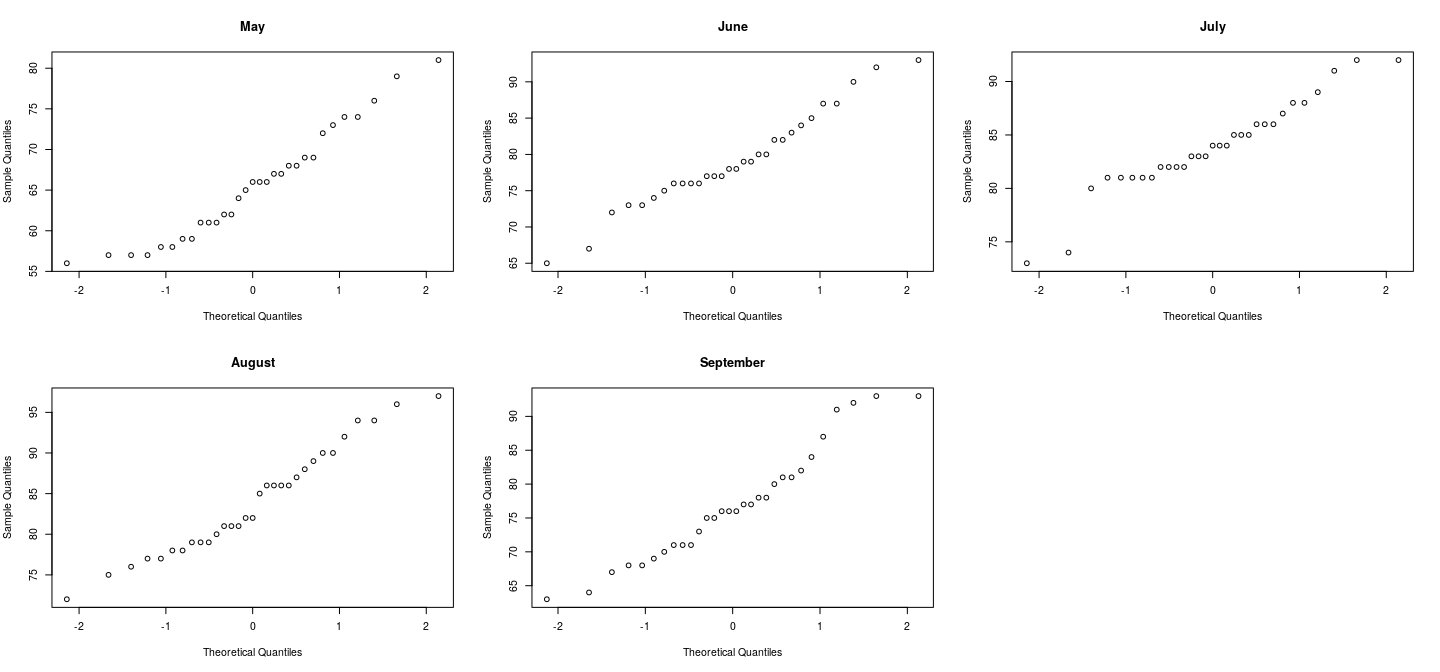
Juxtaposing by splitting figure region
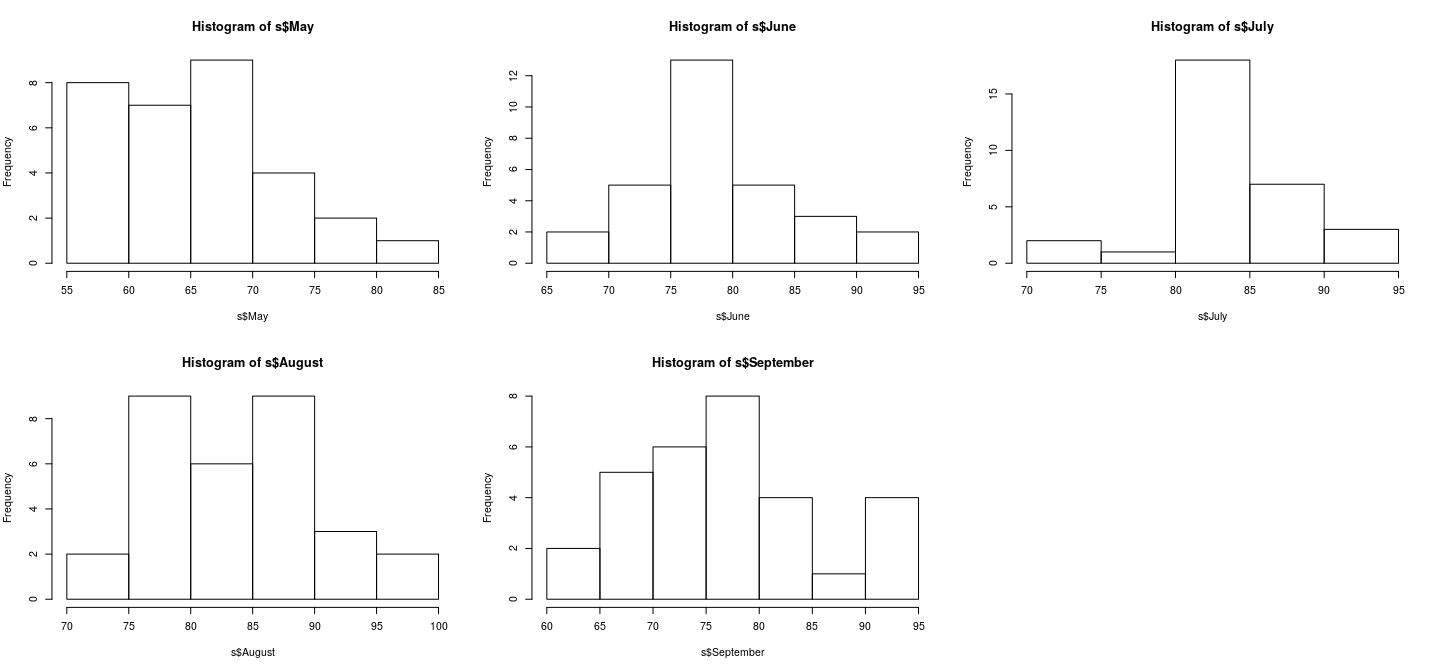
Juxtaposing: better comparison with common scales
par(mfrow = c(2, 3)); r <- range(airquality$Temp)
hist(s$May, xlim = r)
hist(s$June, xlim = r)
hist(s$July, xlim = r)
hist(s$August, xlim = r)
hist(s$September, xlim = r)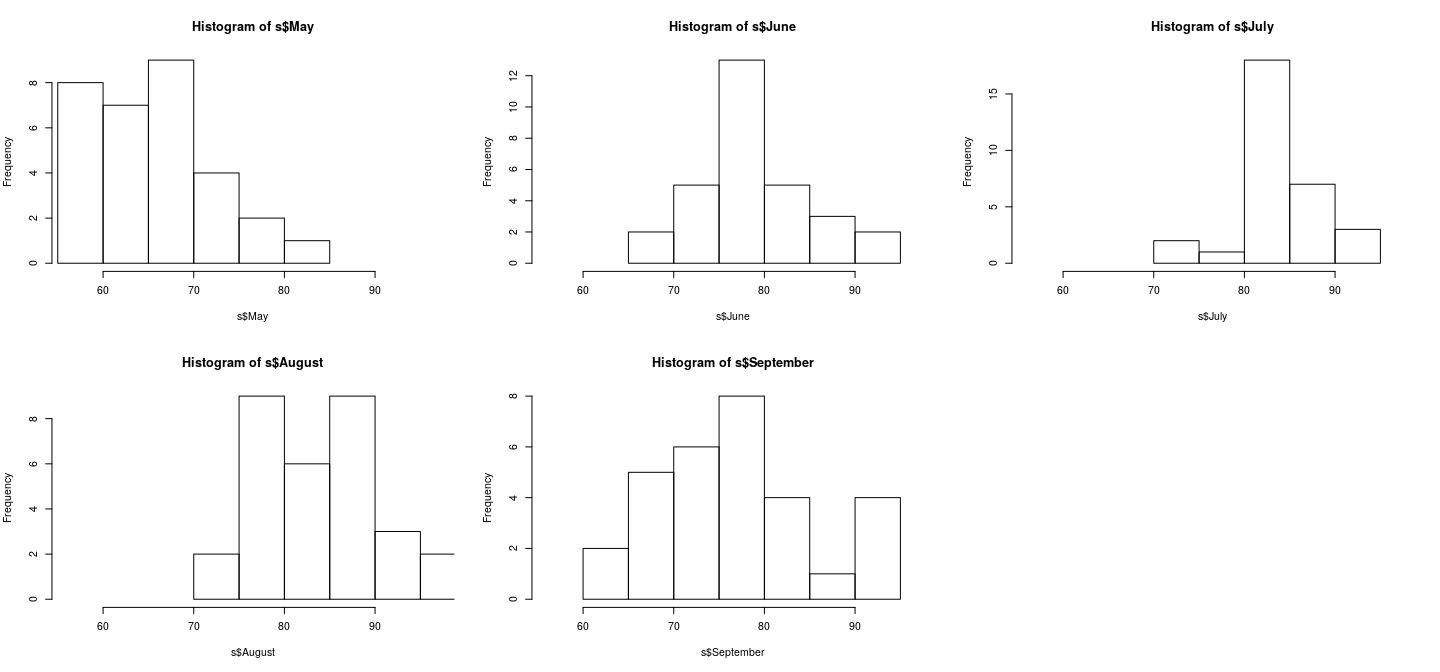
Juxtaposing: better comparison with common scales
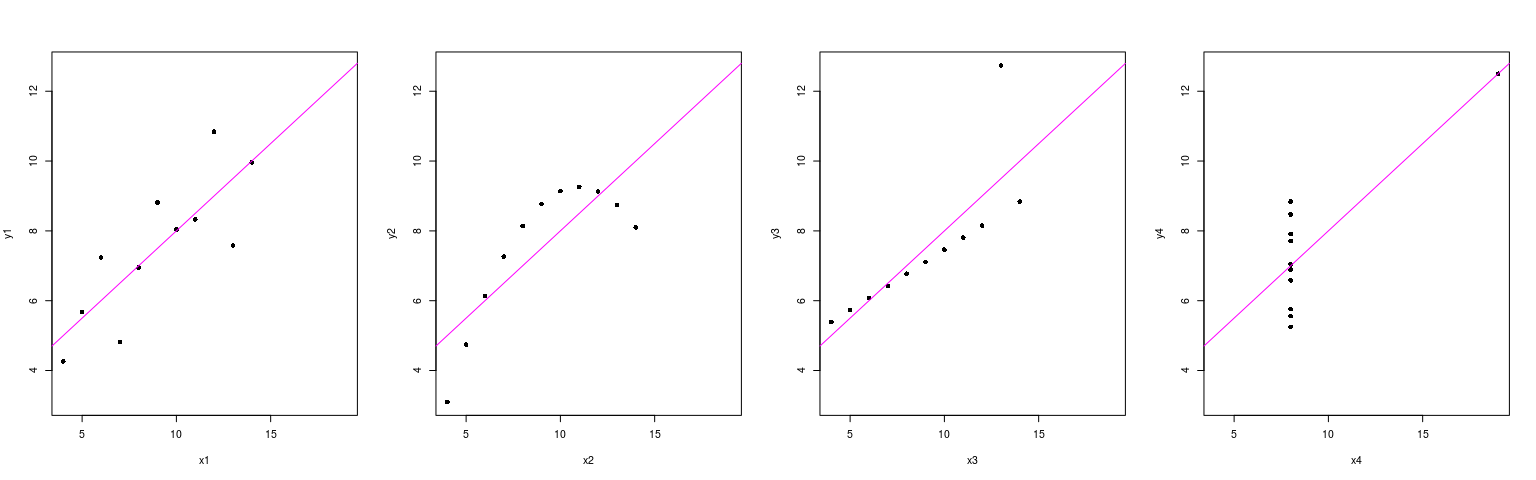
par(mfrow = c(1, 4))
with(anscombe,
{
rx <- range(x1, x2, x3, x4)
ry <- range(y1, y2, y3, y4)
plot(y1 ~ x1, pch = 16, xlim = rx, ylim = ry); abline(lm(y1 ~ x1), col = "magenta")
plot(y2 ~ x2, pch = 16, xlim = rx, ylim = ry); abline(lm(y2 ~ x2), col = "magenta")
plot(y3 ~ x3, pch = 16, xlim = rx, ylim = ry); abline(lm(y3 ~ x3), col = "magenta")
plot(y4 ~ x4, pch = 16, xlim = rx, ylim = ry); abline(lm(y4 ~ x4), col = "magenta")
})
Superposition is better when feasible
dlist <- lapply(s, density, na.rm = TRUE)
dxrng <- range(unlist(lapply(dlist, function(d) d$x)))
dyrng <- range(unlist(lapply(dlist, function(d) d$y)))
plot(dxrng, dyrng, xlab = "Temperature", ylab = "Density")
for (i in seq_along(dlist)) lines(dlist[[i]], col = i)
legend("topright", legend = names(dlist),
lty = 1, col = seq_along(dlist))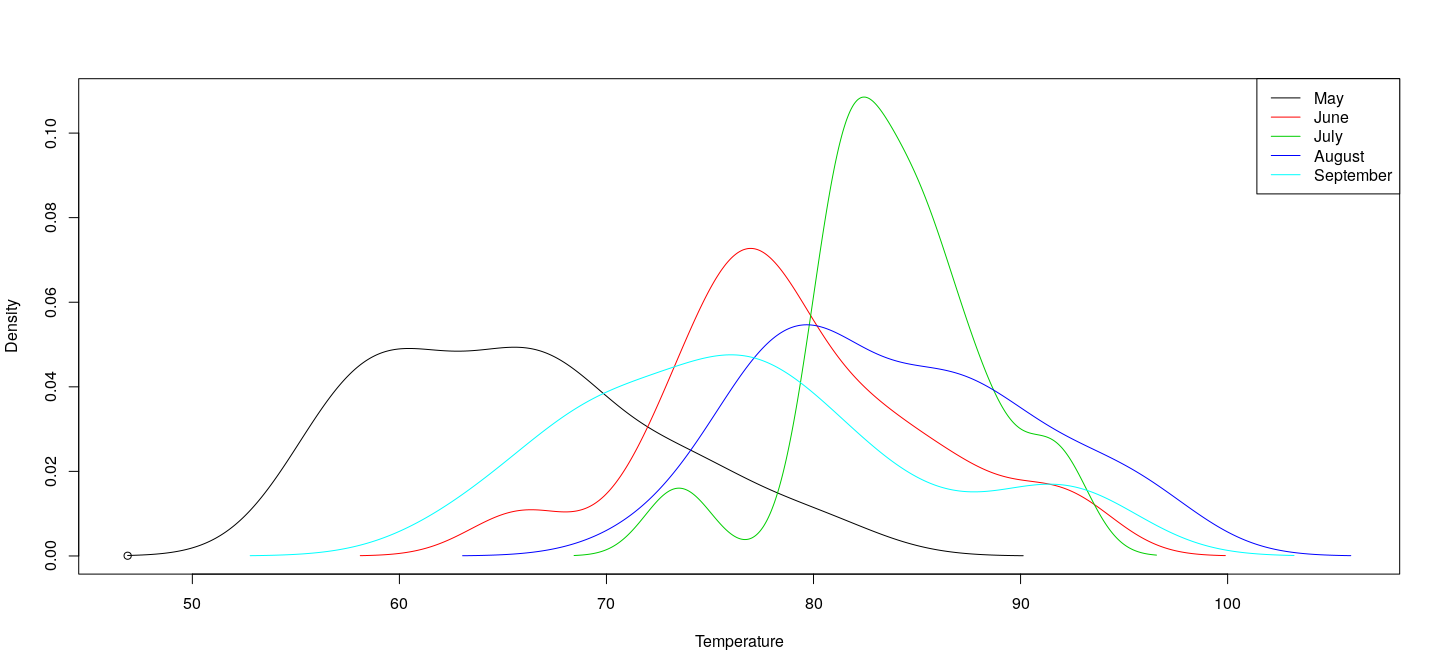
Limitations of traditional graphics
Although not very difficult, these plots are not simple either
The results leave a lot to be desired
Eventually led to the development of alternative systems such as
latticeandggplot2Let’s see some examples for comparison
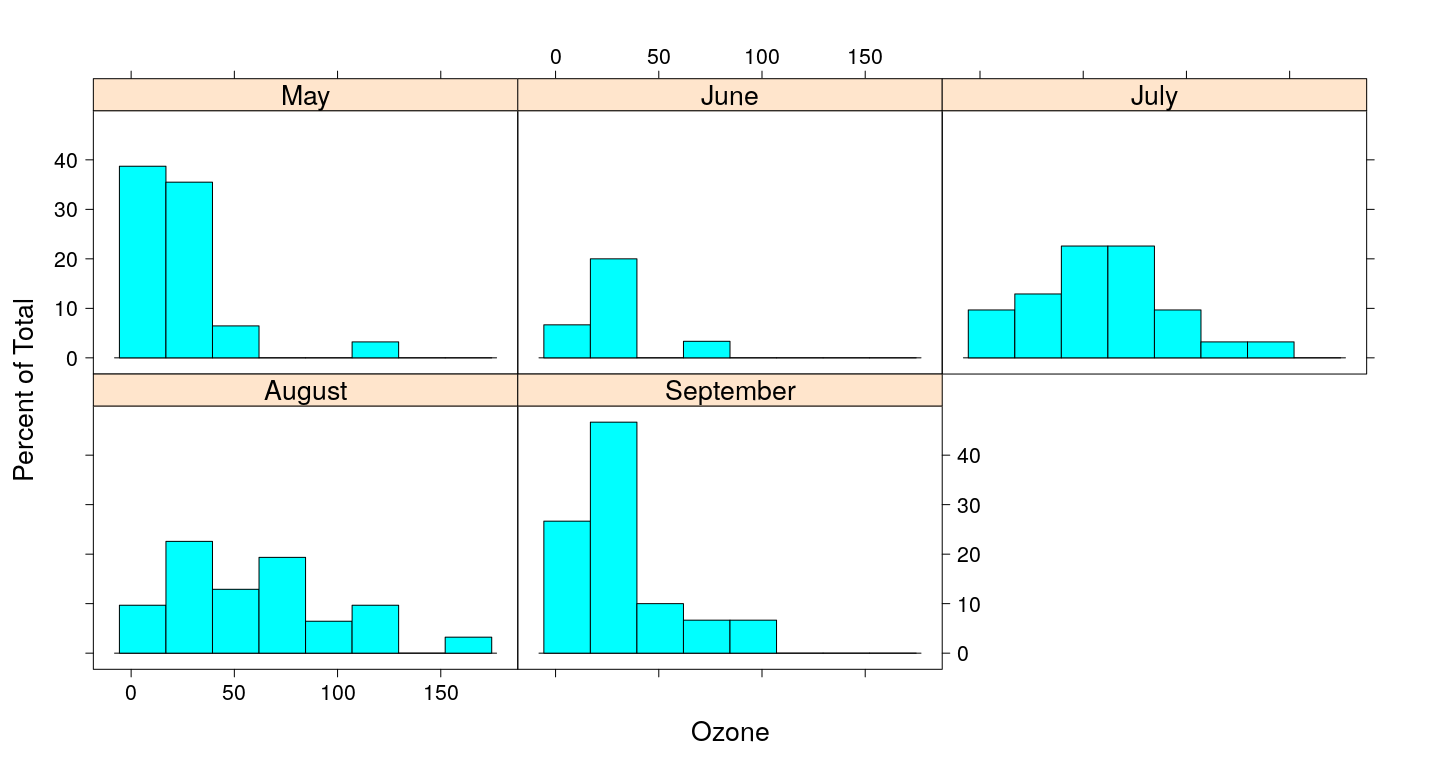
Example of a lattice plot
Example of a lattice plot
Example of a ggplot2 plot
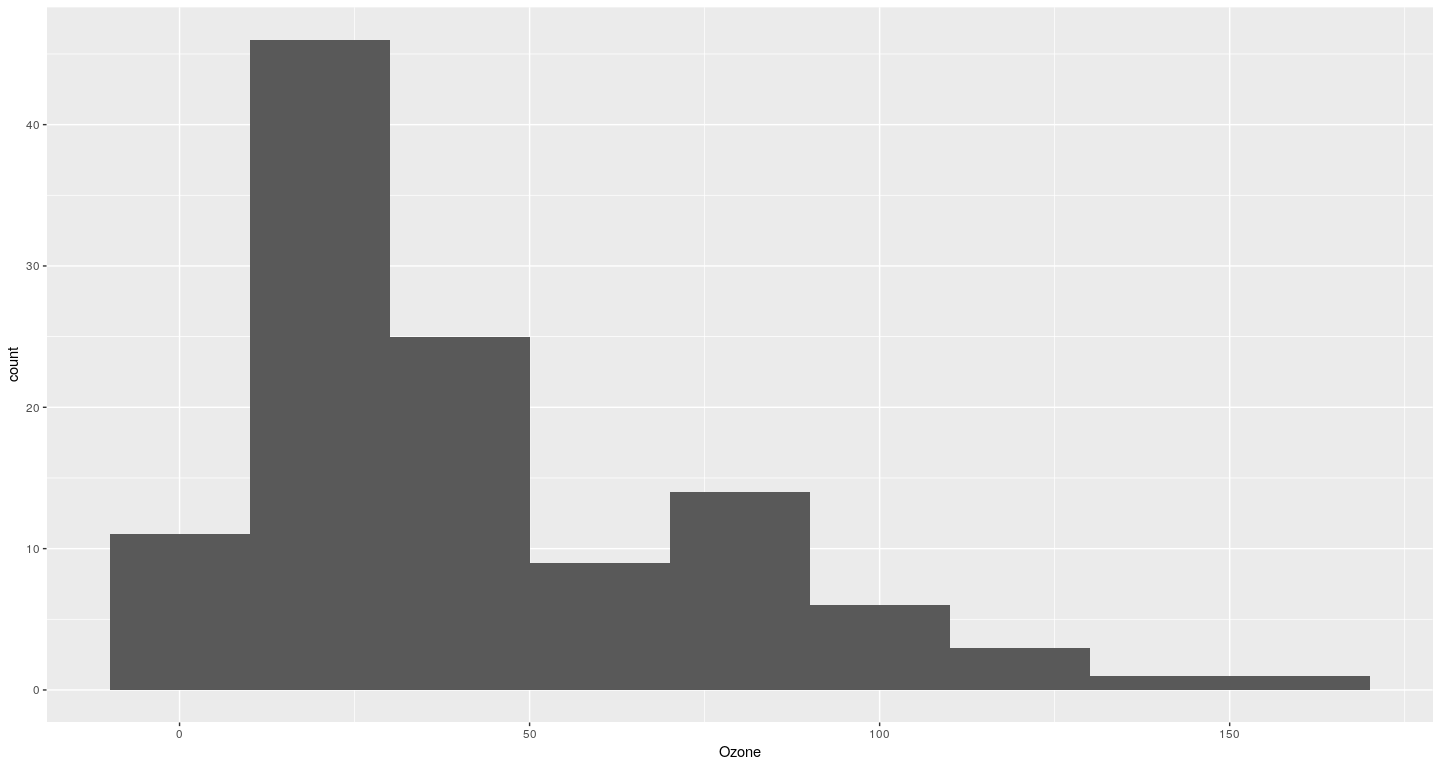
Example of a ggplot2 plot
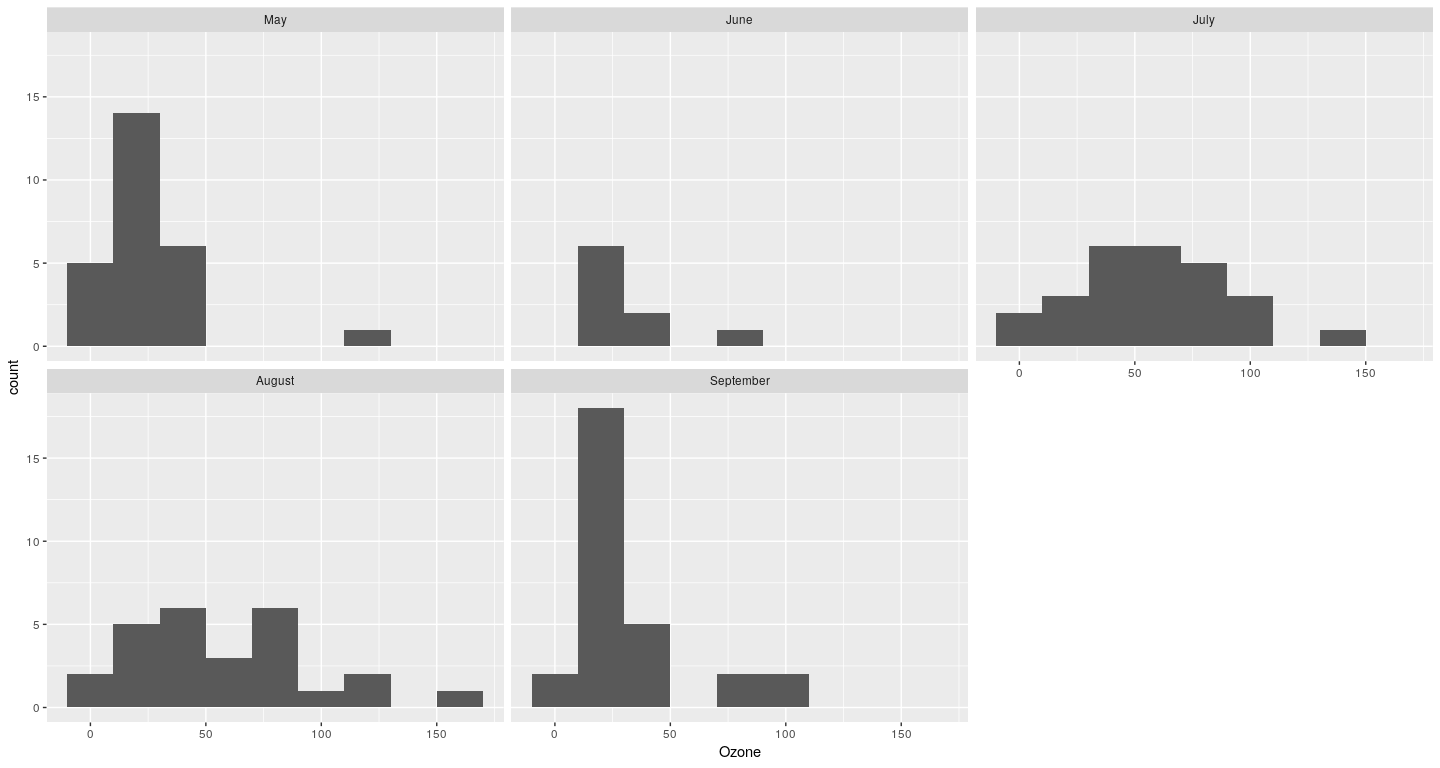
lattice and ggplot2
Both are add-on packages
latticeis based on Trellis graphics in S-PLUSggplot2is based on the “Grammar of Graphics”Two very different philosophical approaches
We will learn about both these in a little more detail
Overview: lattice
Package implementing high-level statistical displays
Philosophically similar to traditional R graphics
Different function for different types of displays (histograms, scatter plots, etc.)
Customization done using low-level functions
Extensively uses formula-data interface
Use as much of the available space as possible
Enable direct comparsion by superposition (grouping) when possible
Encourage comparison when juxtaposing (conditioning):
- use common axes, add common reference objects such as grids.
Example: Scatter plots with xyplot()
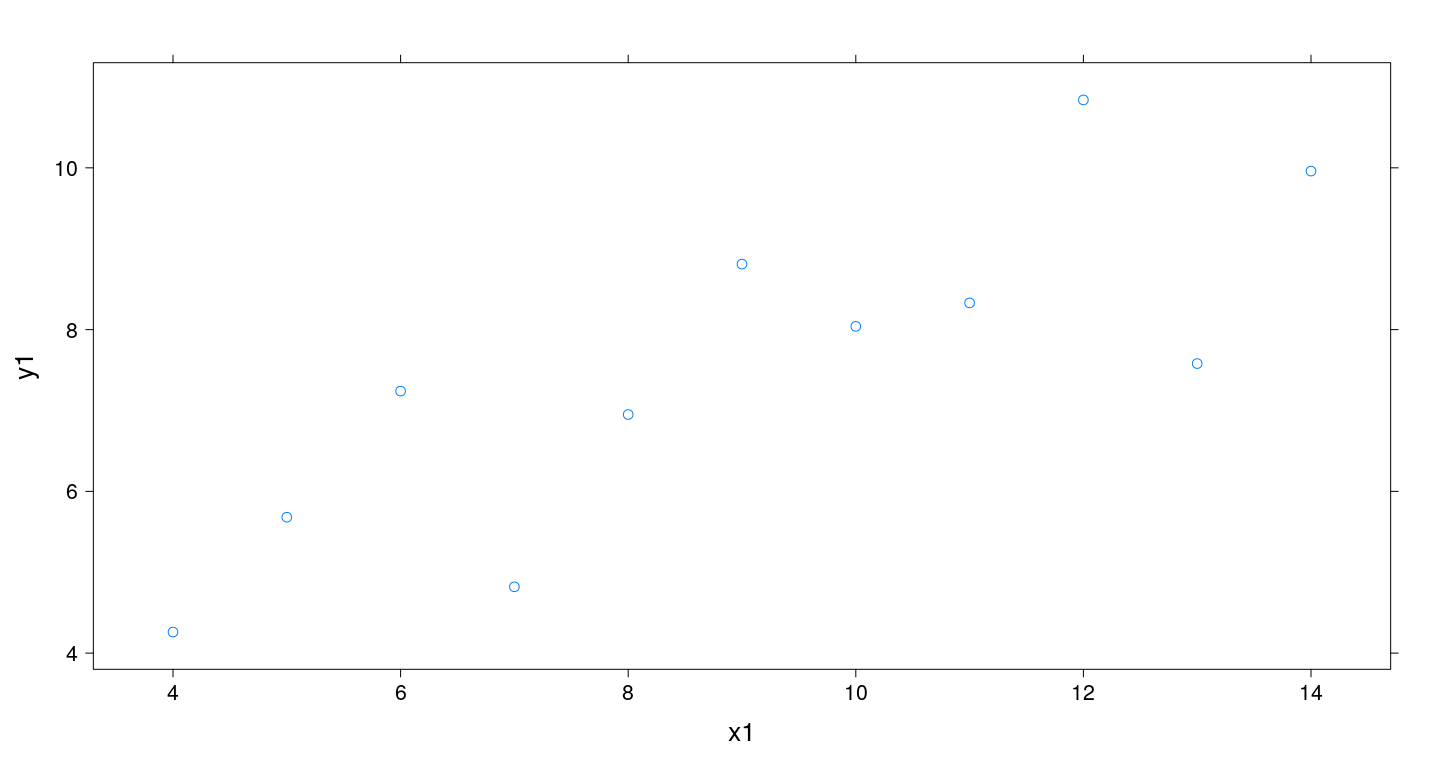
Data must be in suitable form for conditioning
'data.frame': 11 obs. of 8 variables:
$ x1: num 10 8 13 9 11 14 6 4 12 7 ...
$ x2: num 10 8 13 9 11 14 6 4 12 7 ...
$ x3: num 10 8 13 9 11 14 6 4 12 7 ...
$ x4: num 8 8 8 8 8 8 8 19 8 8 ...
$ y1: num 8.04 6.95 7.58 8.81 8.33 ...
$ y2: num 9.14 8.14 8.74 8.77 9.26 8.1 6.13 3.1 9.13 7.26 ...
$ y3: num 7.46 6.77 12.74 7.11 7.81 ...
$ y4: num 6.58 5.76 7.71 8.84 8.47 7.04 5.25 12.5 5.56 7.91 ...anscombe.long <-
with(anscombe,
data.frame(x = c(x1, x2, x3, x4),
y = c(y1, y2, y3, y4),
which = factor(rep(1:4, each = nrow(anscombe)))))
str(anscombe.long) # OK'data.frame': 44 obs. of 3 variables:
$ x : num 10 8 13 9 11 14 6 4 12 7 ...
$ y : num 8.04 6.95 7.58 8.81 8.33 ...
$ which: Factor w/ 4 levels "1","2","3","4": 1 1 1 1 1 1 1 1 1 1 ...Example: conditioning
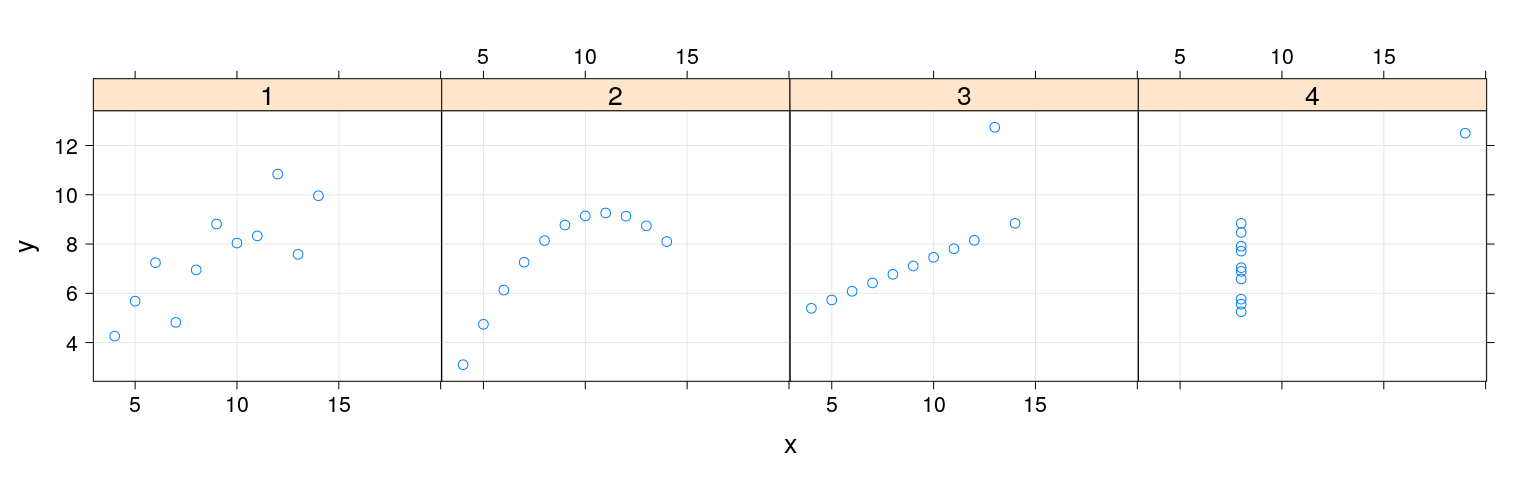
Here x and y are “primary variables”, which is a “conditioning variable”.
Customization
How can we add regression lines as before?
There is not one but four lines to add
- The solution in
latticeis to define a procedure to display data:
Customization
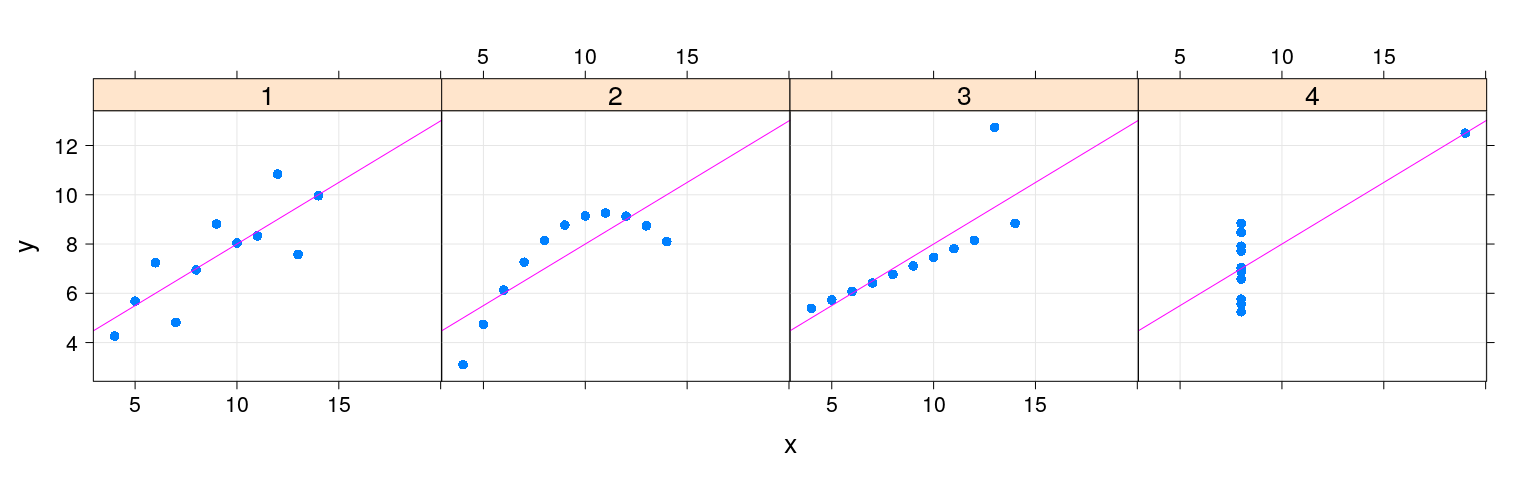
Overview: lattice
Whole display created in one step; “work-in-progress” model does not work
Each high-level plot has a default display
Variables play different roles: primary, conditioning, grouping (superposition)
Can be customized by a user-supplied “panel” function
In fact, many other aspects can be customized: axis annotation, strips, legends
- We will see some more examples of different high-level functions before moving on to
ggplot2
Histogram
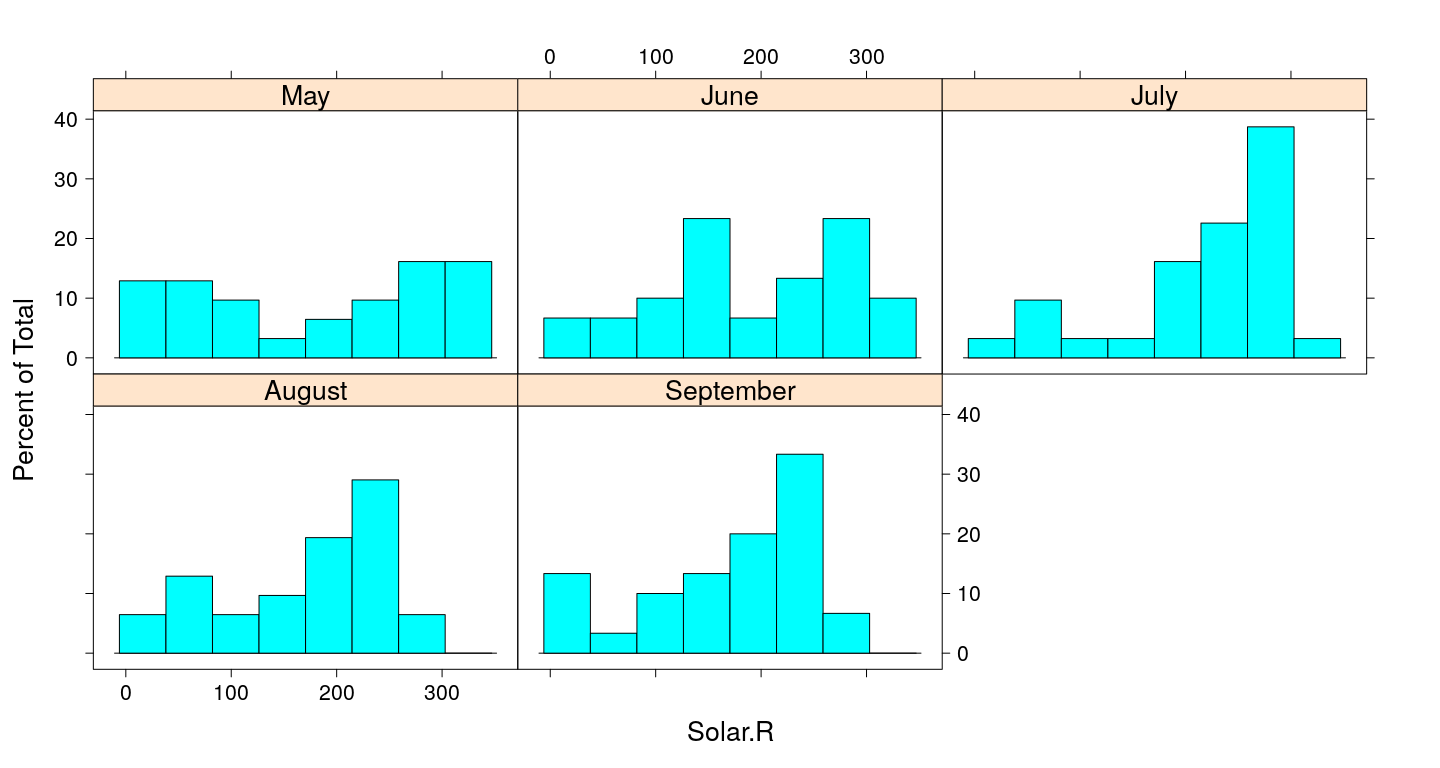
Kernel density plots
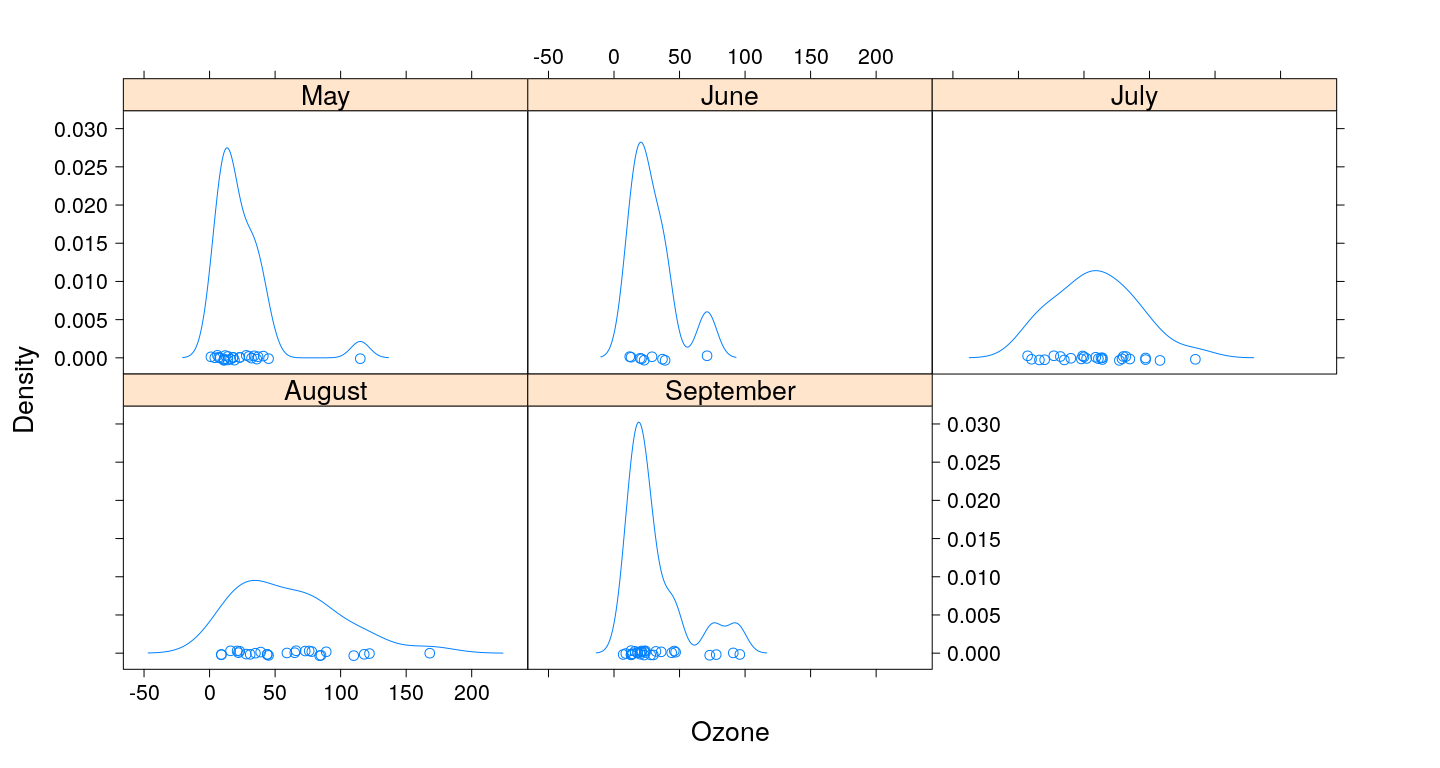
Kernel density plots (with grouping)
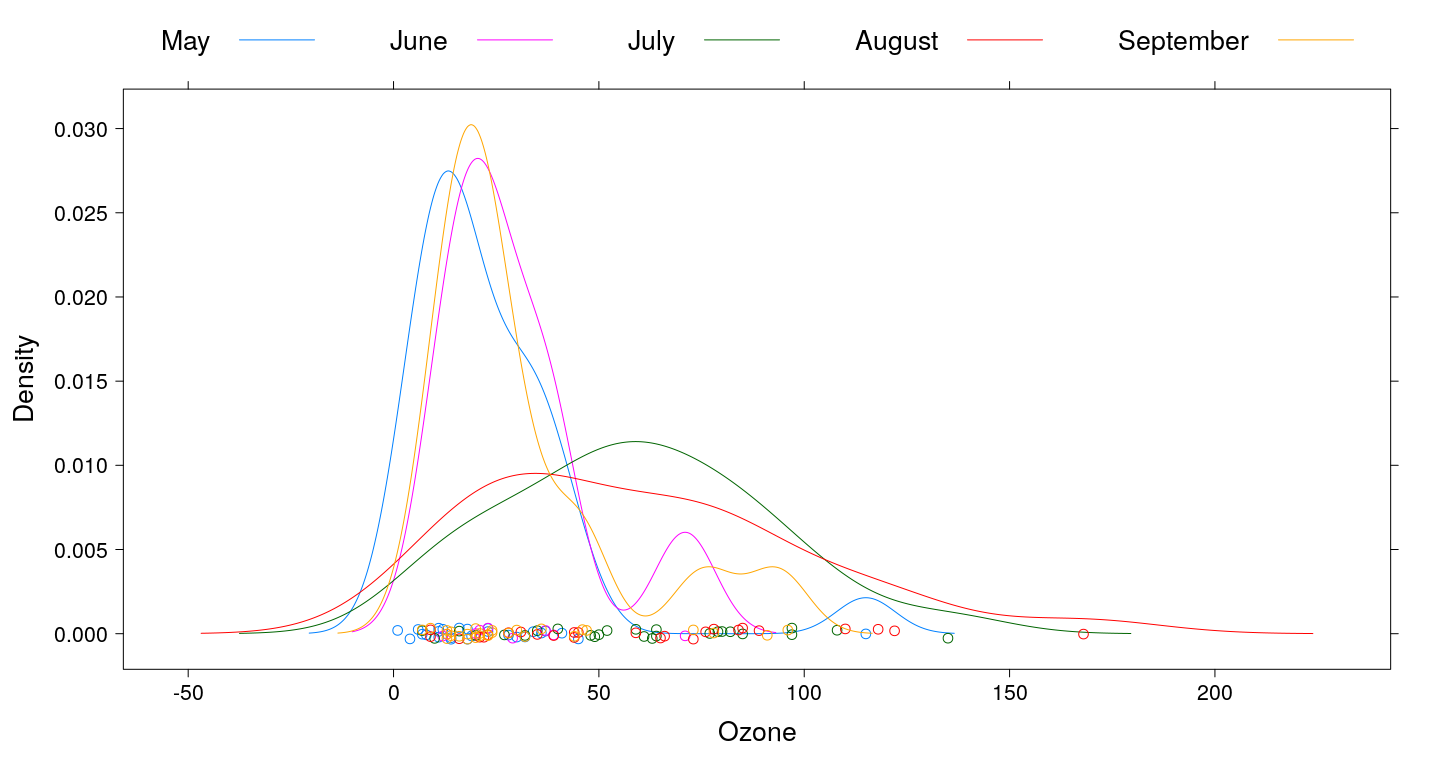
Q-Q plots
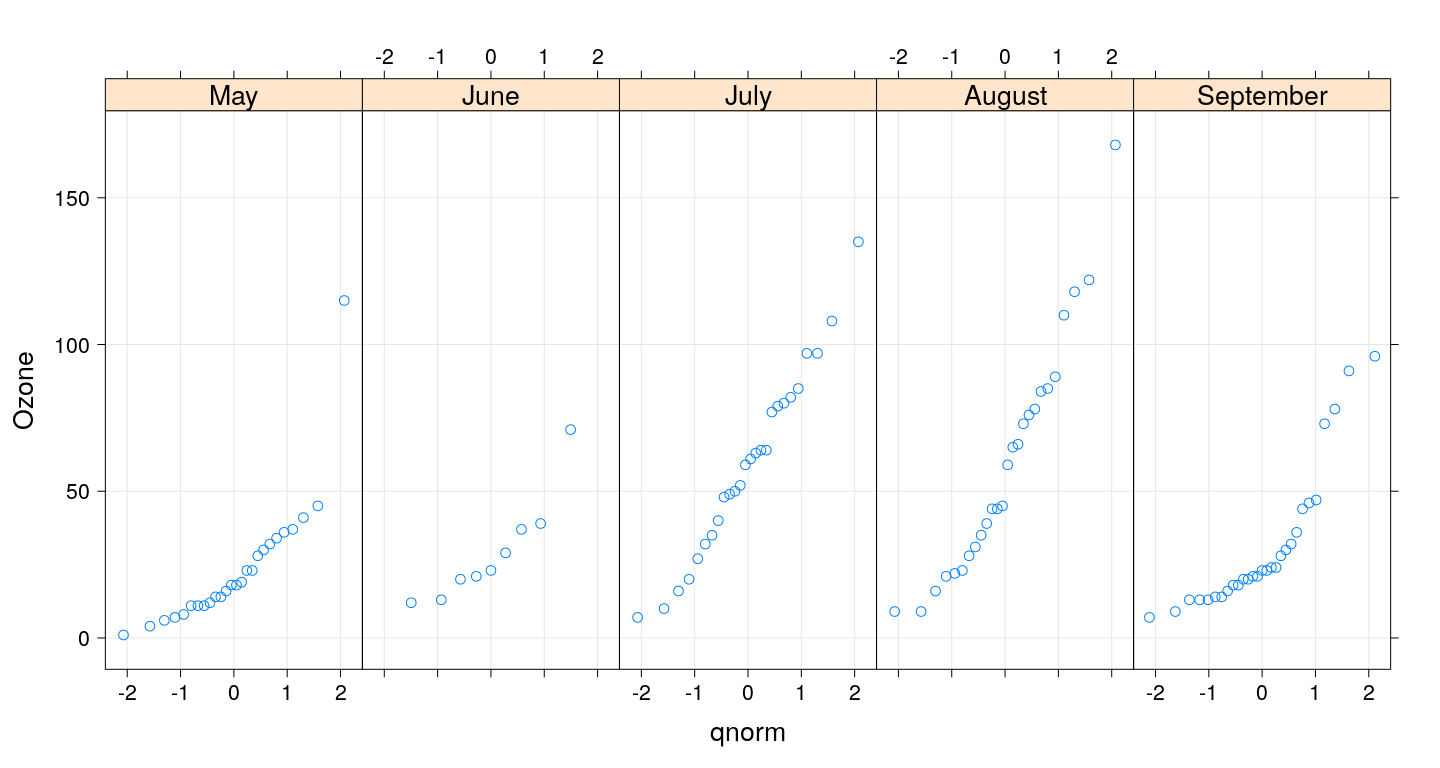
Box and whisker plots
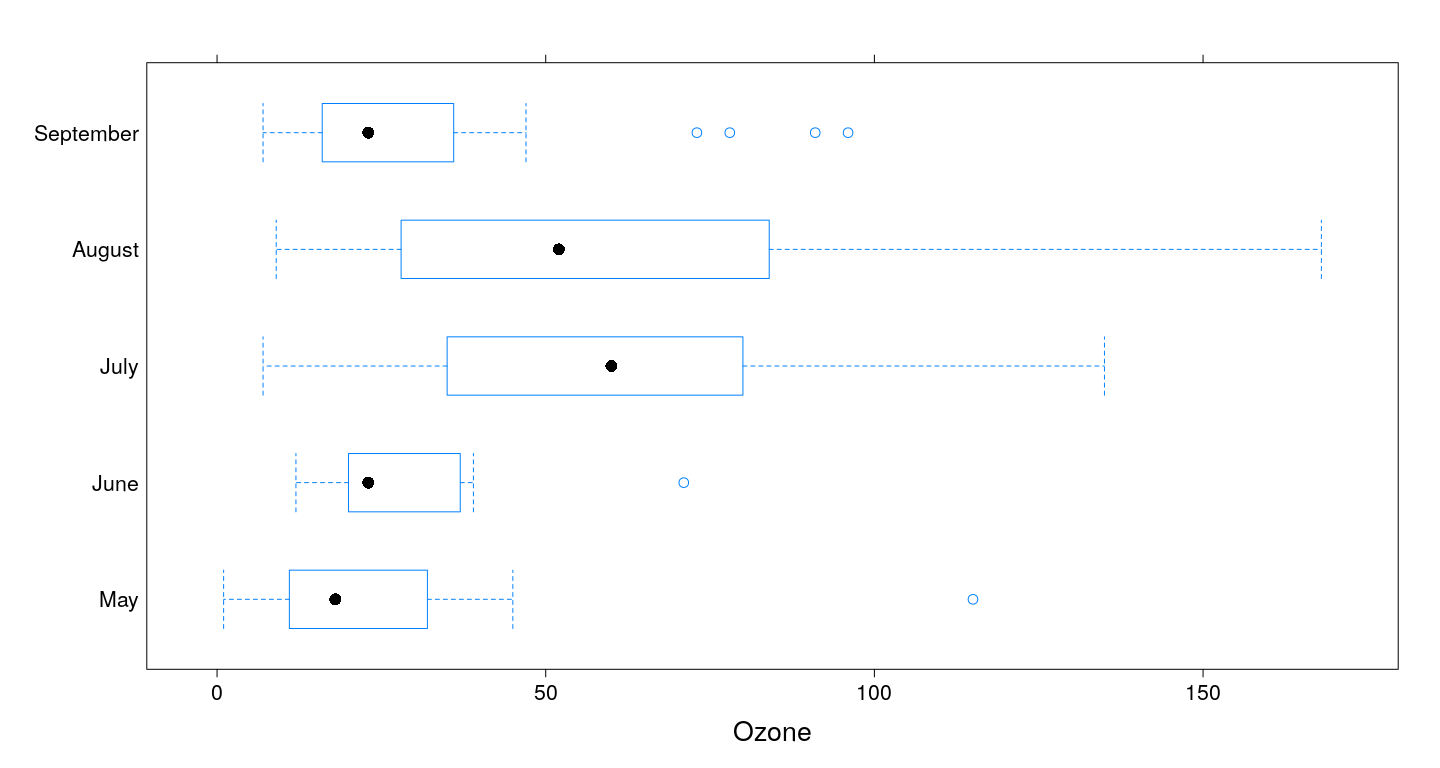
Multiple primary variables / different scales
bwplot(fmonth ~ Ozone + Solar.R, airquality, outer = TRUE,
between = list(x = 1),
scales = list(x = "free"), xlab = NULL)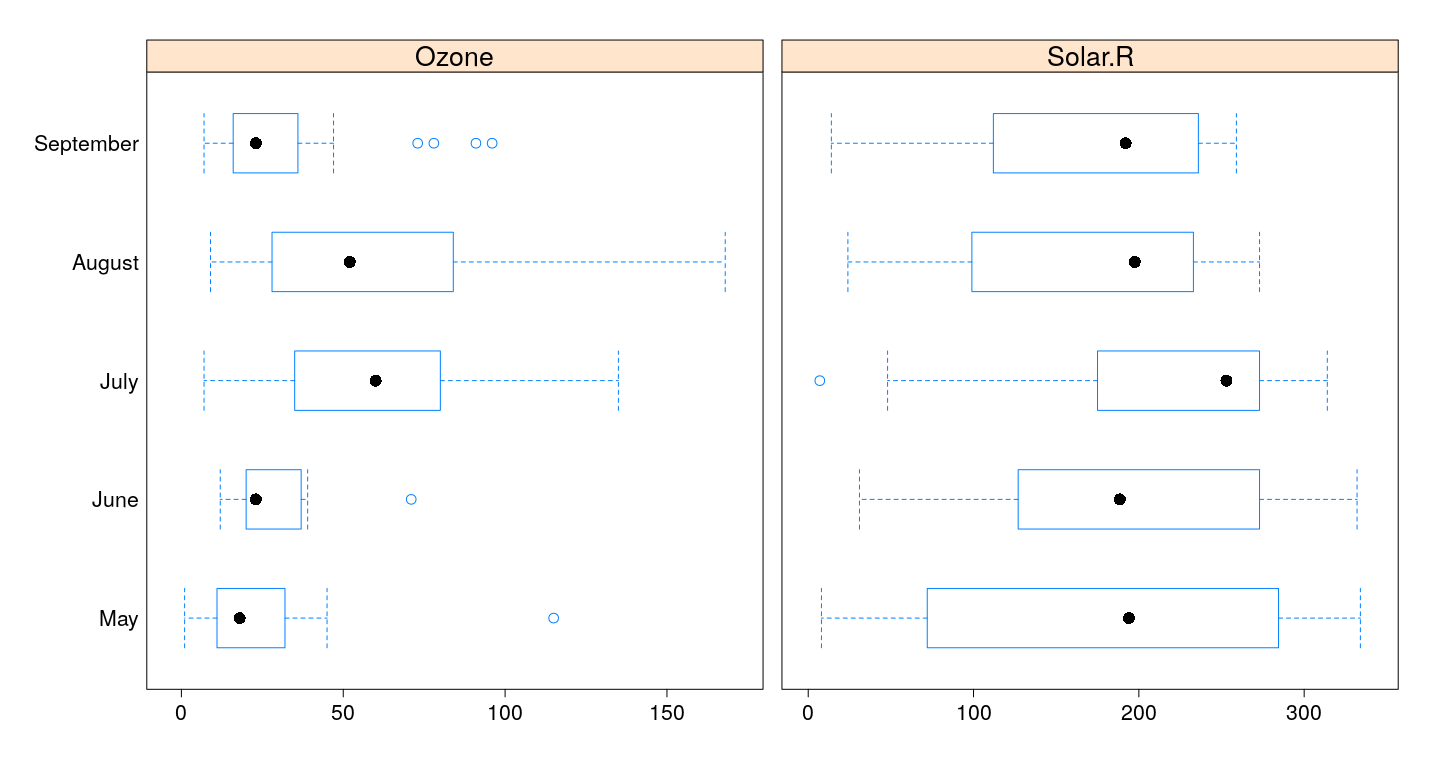
Violin plots - alternative display function
bwplot(fmonth ~ Ozone + Solar.R, airquality, outer = TRUE,
between = list(x = 1), panel = panel.violin,
scales = list(x = "free"), xlab = NULL)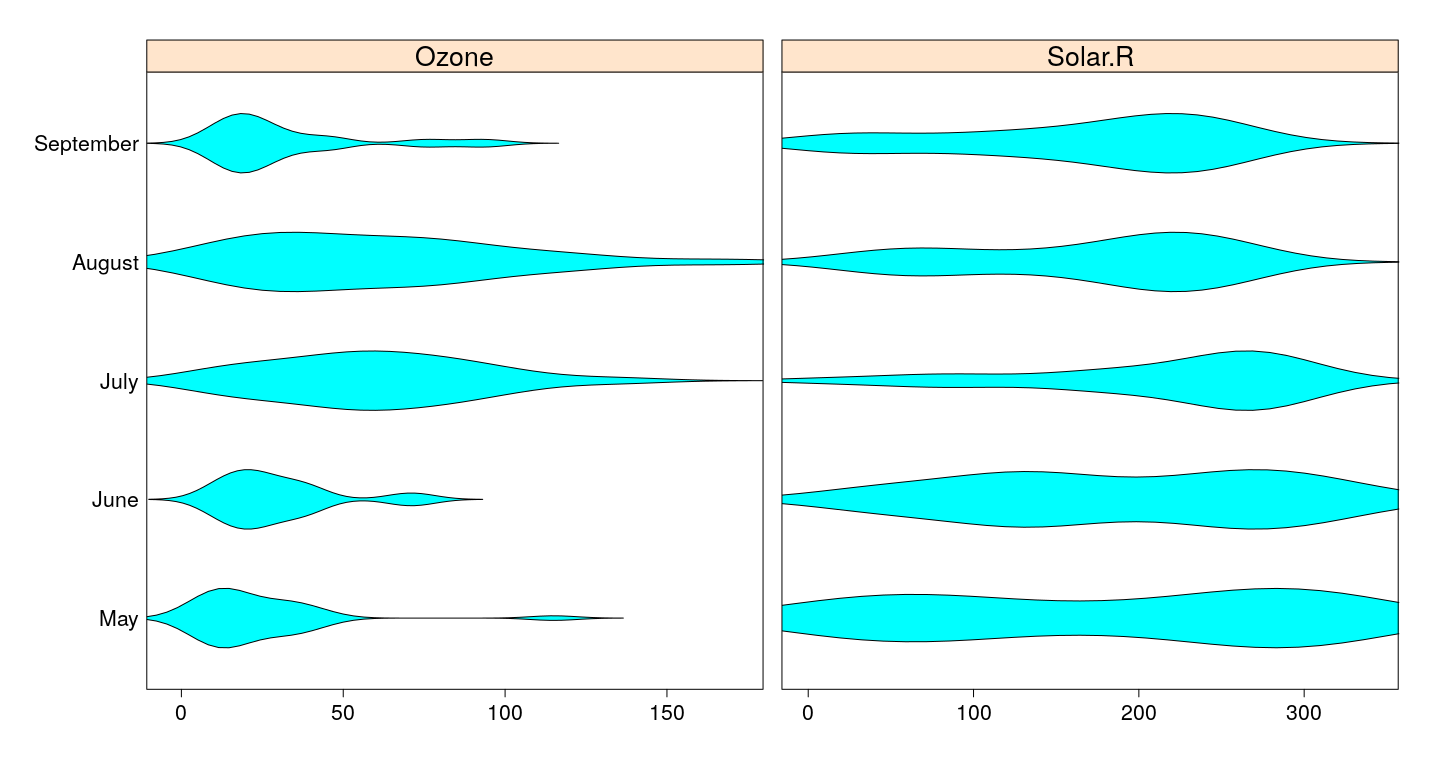
Tabular data
Some common displays are designed for tabular data: bar chart, dot plot, pie chart
Data are typically counts or rates obtained by cross classification by multiple factors
Rural Male Rural Female Urban Male Urban Female
50-54 11.7 8.7 15.4 8.4
55-59 18.1 11.7 24.3 13.6
60-64 26.9 20.3 37.0 19.3
65-69 41.0 30.9 54.6 35.1
70-74 66.0 54.3 71.1 50.0VADeathsDF <- as.data.frame.table(VADeaths, responseName = "Rate")
str(VADeathsDF) # Better format for graphics functions'data.frame': 20 obs. of 3 variables:
$ Var1: Factor w/ 5 levels "50-54","55-59",..: 1 2 3 4 5 1 2 3 4 5 ...
$ Var2: Factor w/ 4 levels "Rural Male","Rural Female",..: 1 1 1 1 1 2 2 2 2 2 ...
$ Rate: num 11.7 18.1 26.9 41 66 8.7 11.7 20.3 30.9 54.3 ...Bar chart - traditional graphics
par(mfrow = c(1, 2))
barplot(t(VADeaths))
barplot(VADeaths, beside=TRUE, horiz=TRUE,
legend.text=TRUE, xlim = c(0, 100))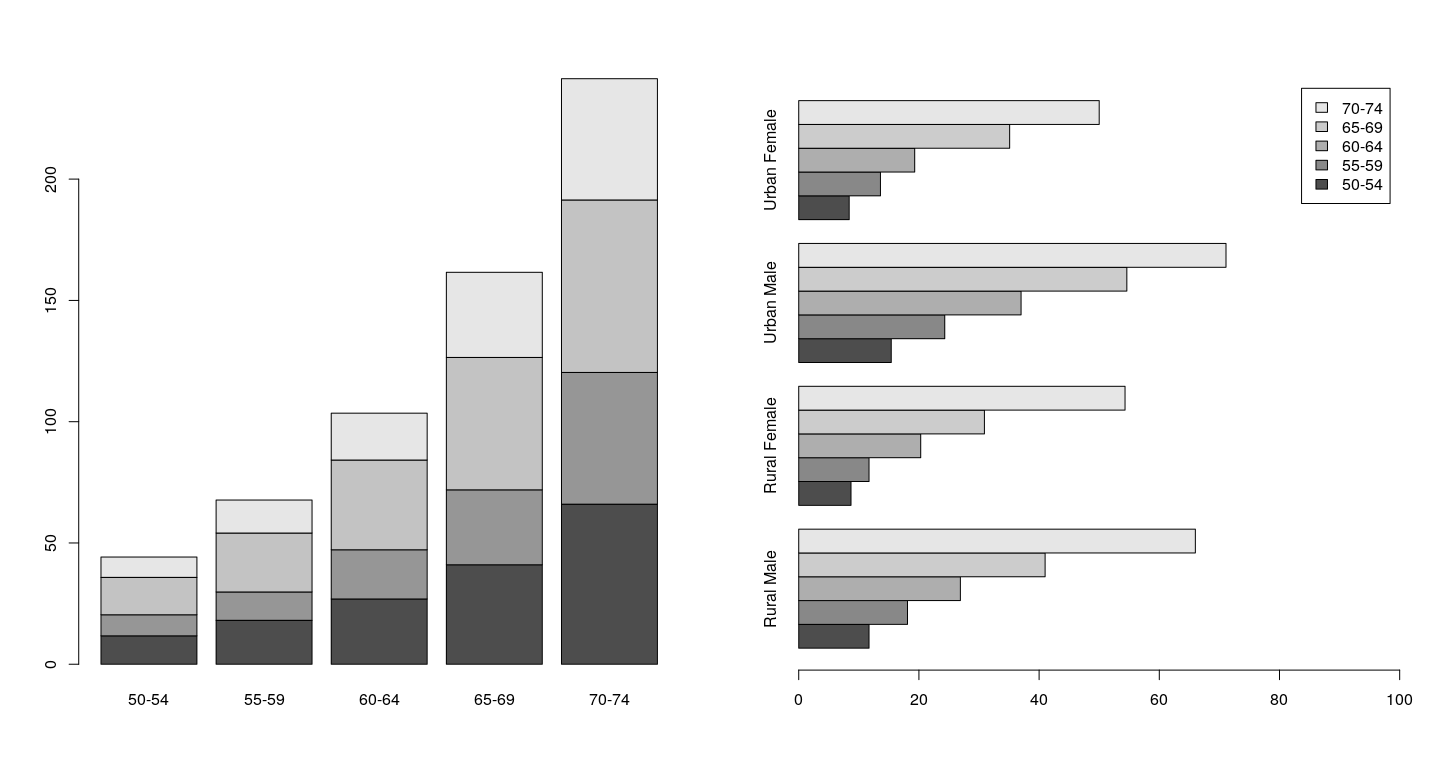
Bar chart - lattice
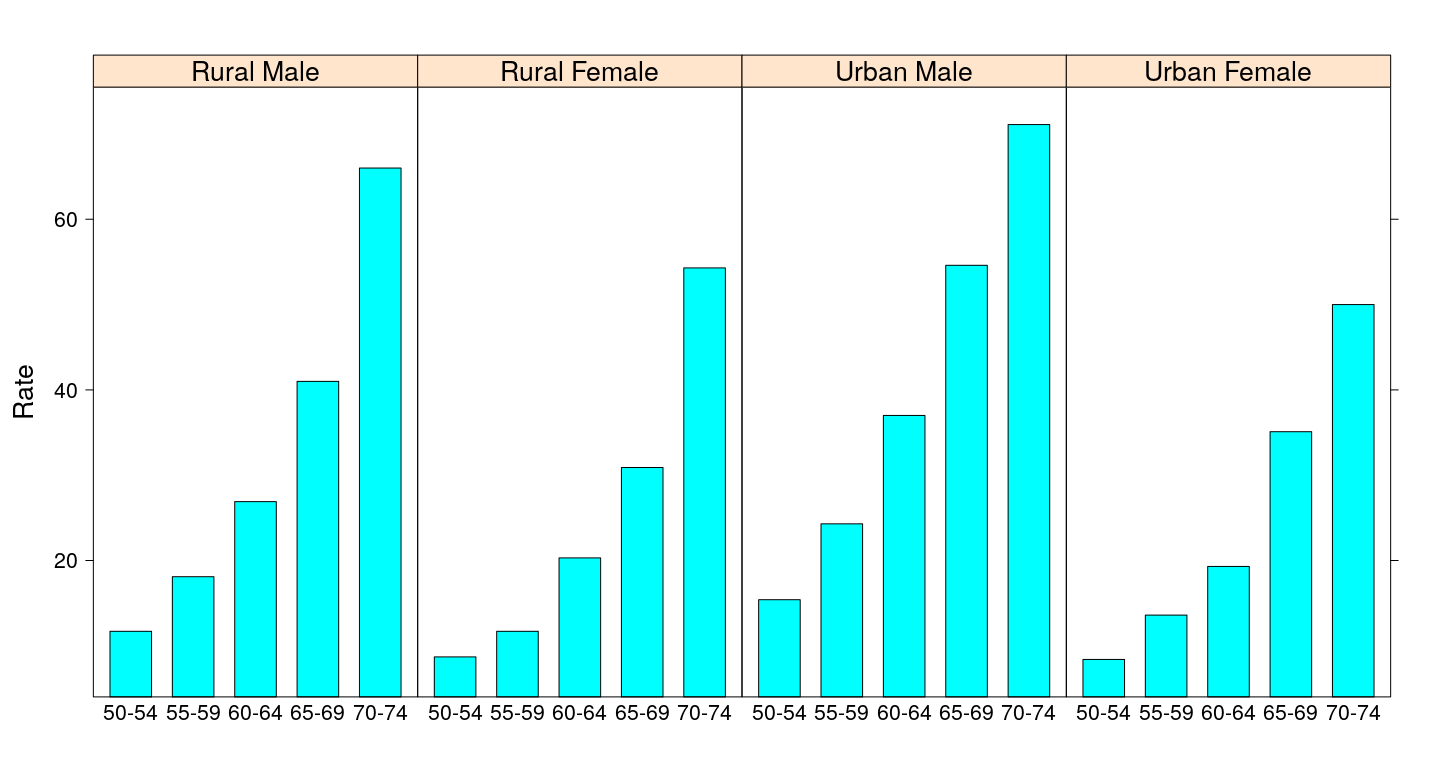
Bar chart - without misleading heights
barchart(Rate ~ Var1 | Var2, VADeathsDF, layout = c(4, 1), origin = 0,
scales = list(x = list(rot = 45)))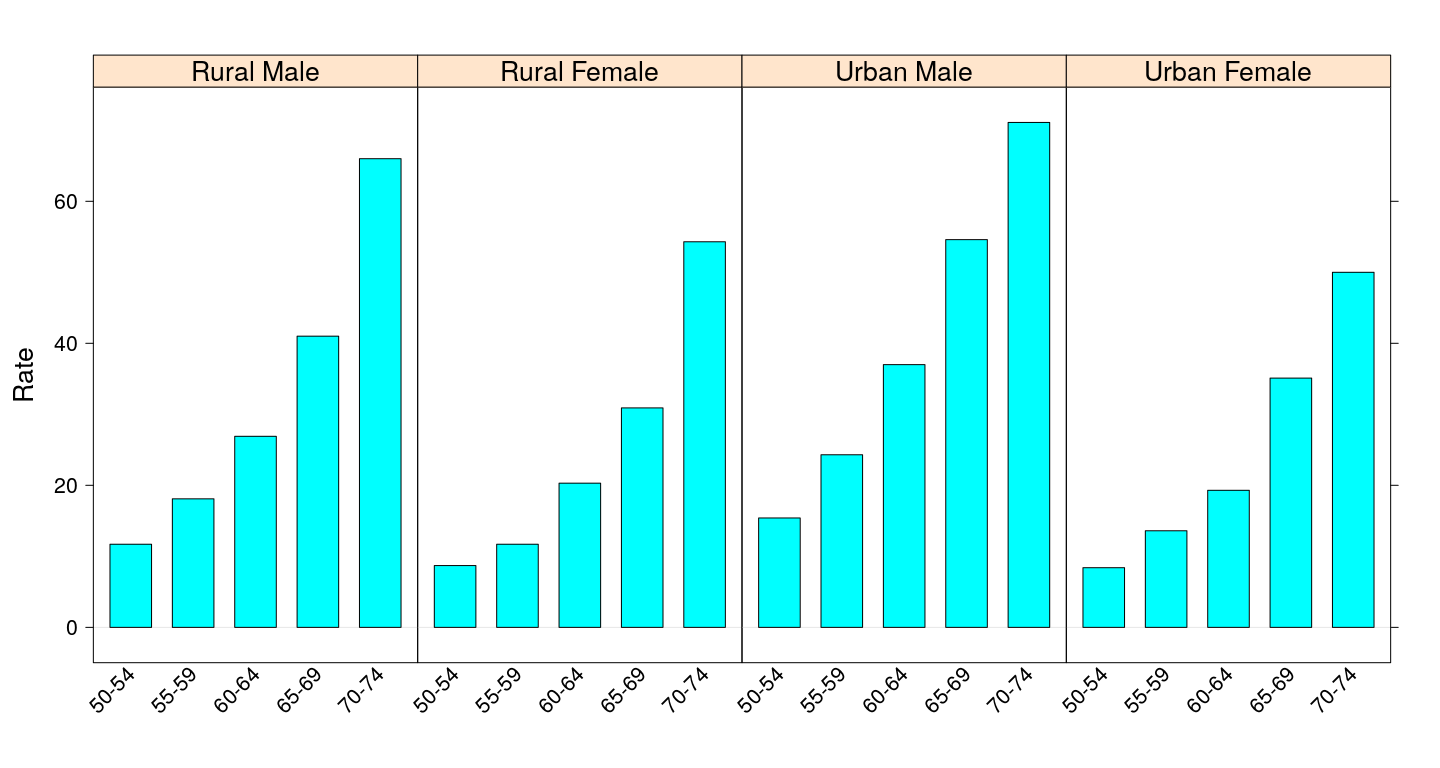
Dot plot
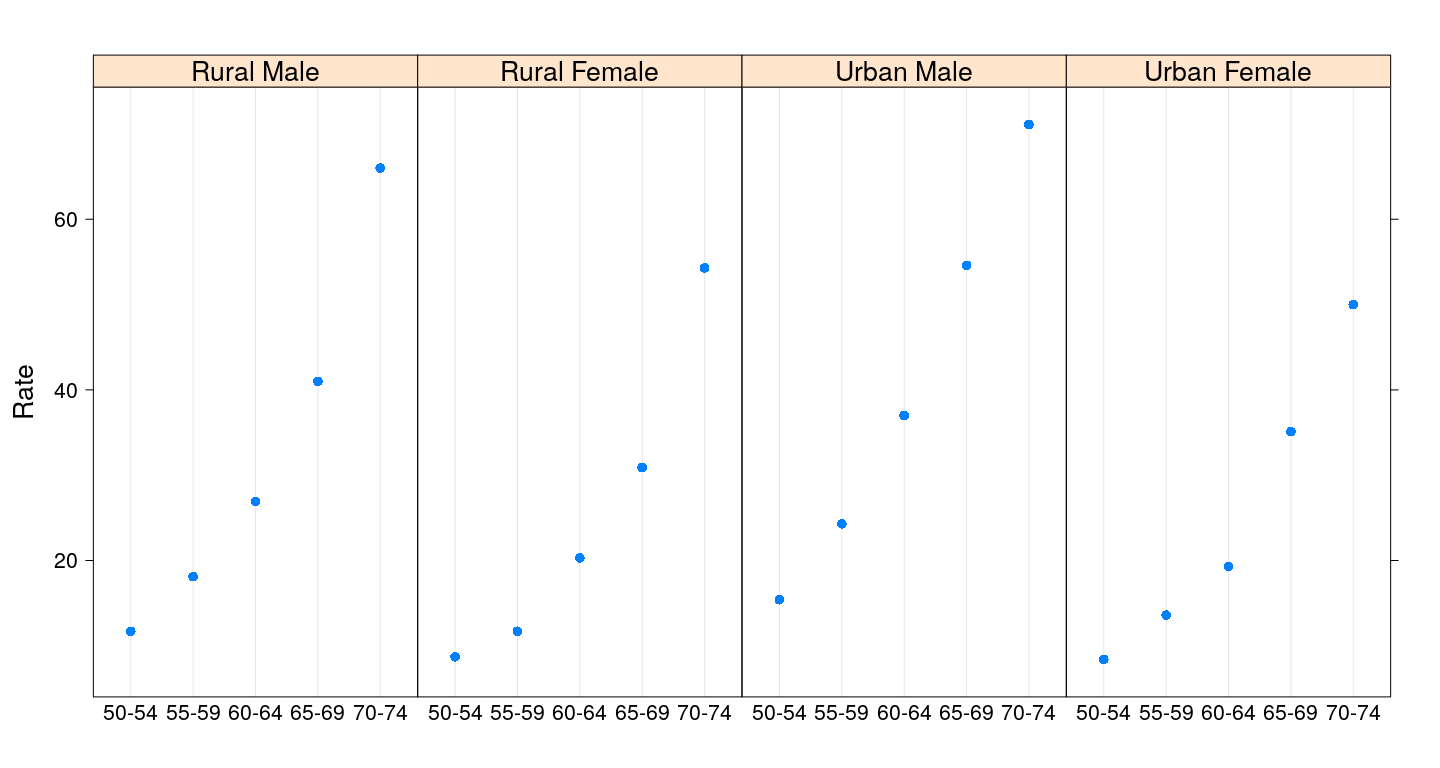
Dot plot with grouping
dotplot(Rate ~ Var1, VADeathsDF, groups = Var2, type = "o",
auto.key = list(columns = 2, points = TRUE, lines = TRUE))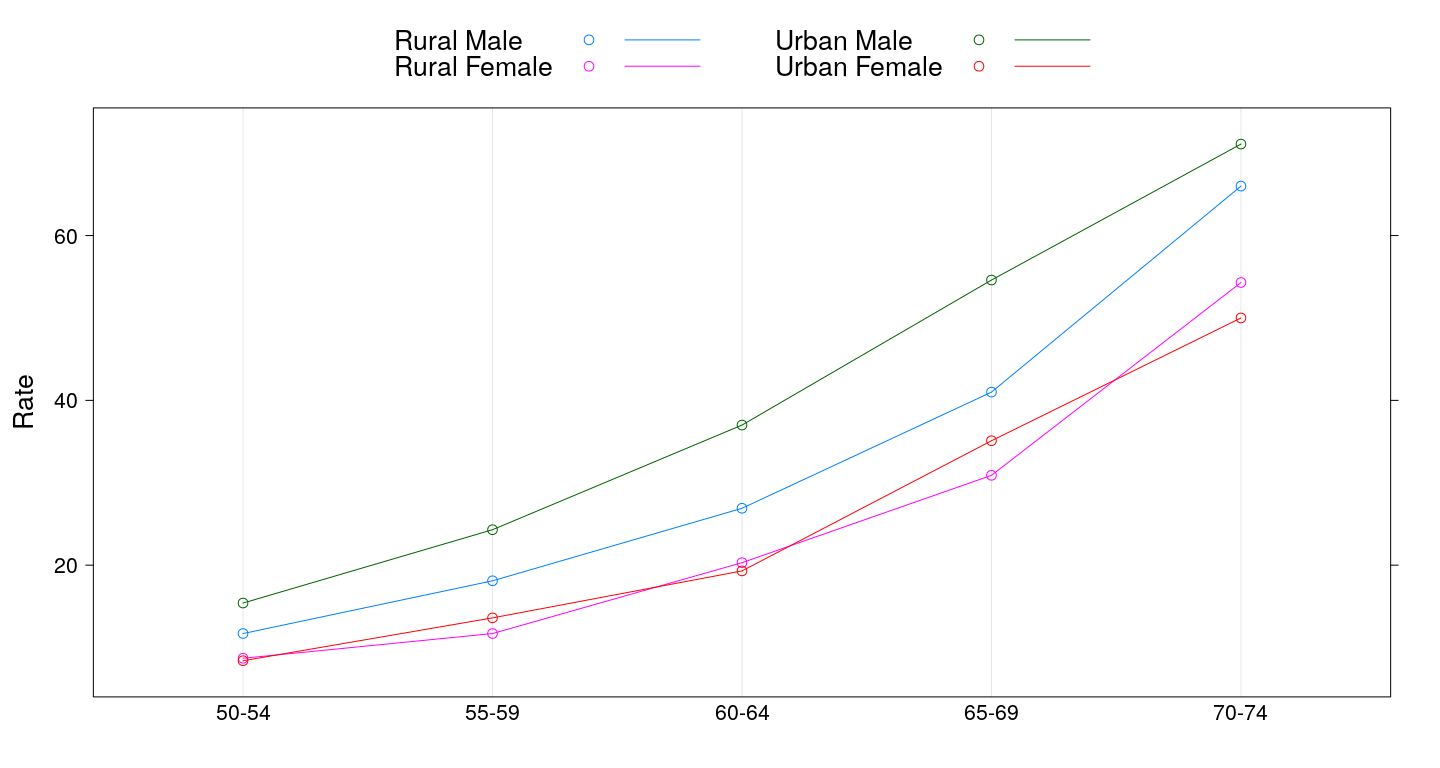
3-D scatter plots
cloud(depth ~ lat * long, data = quakes,
zlim = rev(range(quakes$depth)),
screen = list(z = 105, x = -70), panel.aspect = 0.75,
xlab = "Longitude", ylab = "Latitude", zlab = "Depth")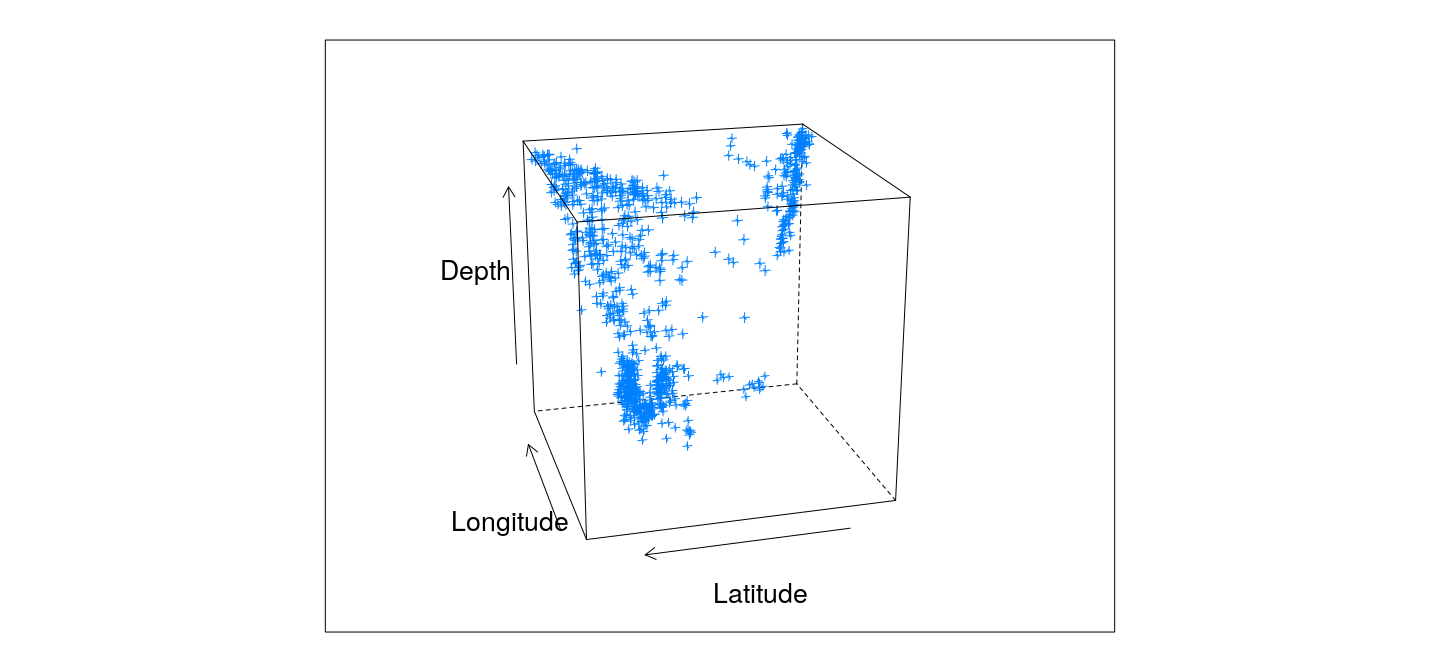
3-D surface plots
wireframe(Freq ~ Var1 + Var2, data = as.data.frame.table(volcano),
shade = TRUE, aspect = c(61/87, 0.5))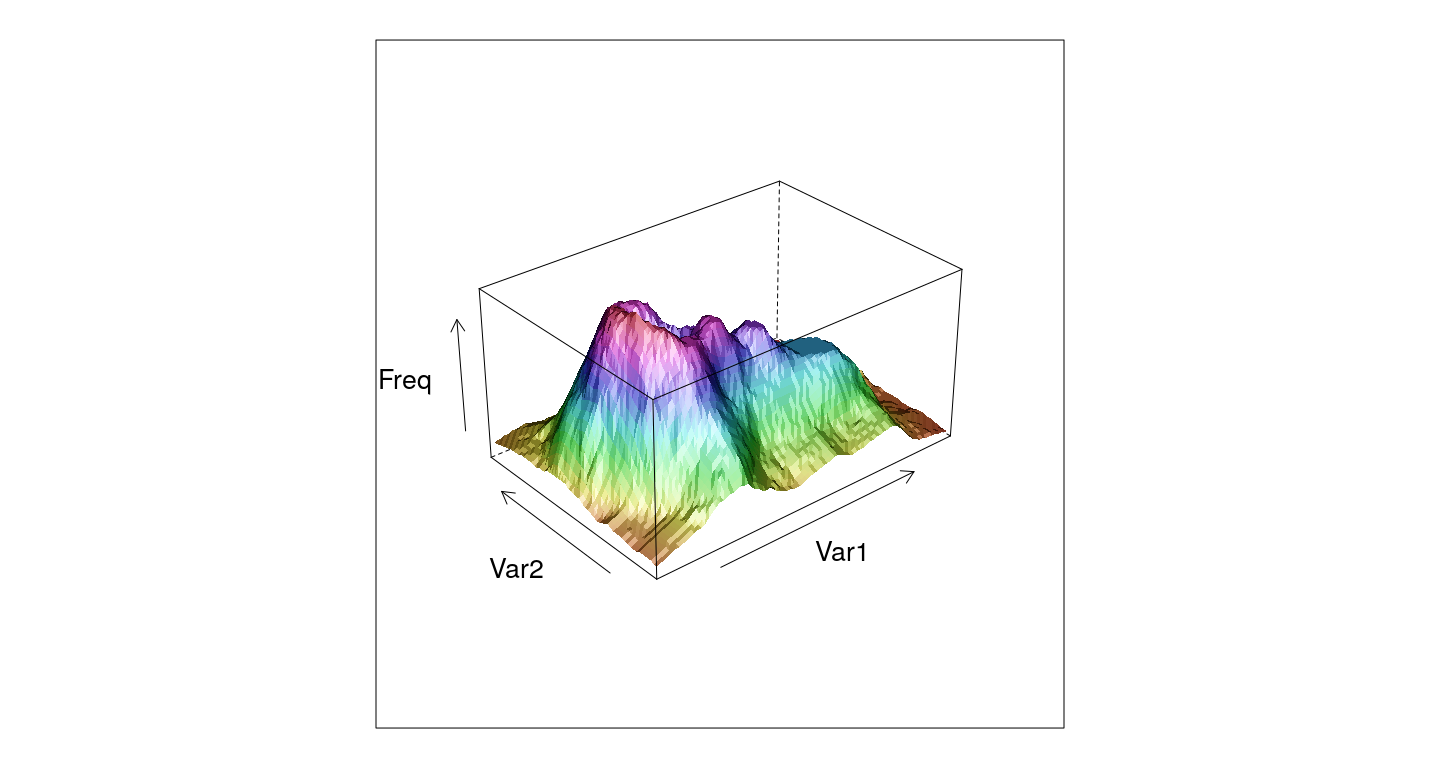
Summary
Separate function for each display type
Display can be customized
Many other advanced features - see manual (start with
package?lattice)
GGplot
Traditional graphics and
latticeare both procedural in their approachThe “grammar of graphics” takes a declarative approach
The user describes a plot using a layered grammar
A plot is composed by “adding” various components
Has one or more layers, each associated with a dataset
Rather than using predefined designs, user describes each layer
Components of the grammar
- Aesthetic mappings that map data values to some aspect of the displayed graph, e.g.,
- coordinate positions
- color, shape, size
- group, etc.
- Geometric type that is used to render the mapped data, e.g.,
- points, lines, polygons, etc.
- something more complex such as a box-and-whisker plot
- Statistical transformations that are applied to the data, e.g.,
- binning for histograms
- computation of kernel density estimates
- Scales that give a visual indication of the aesthetic mappings, e.g.,
- axis annotation for position mapping,
- legends for mapping to color, size, etc.
- Faceting (conditioning in
lattice) to produce small multiples.
Example: scatter plot
- A scatter plot needs a dataset, an x-variable, and a y-variable
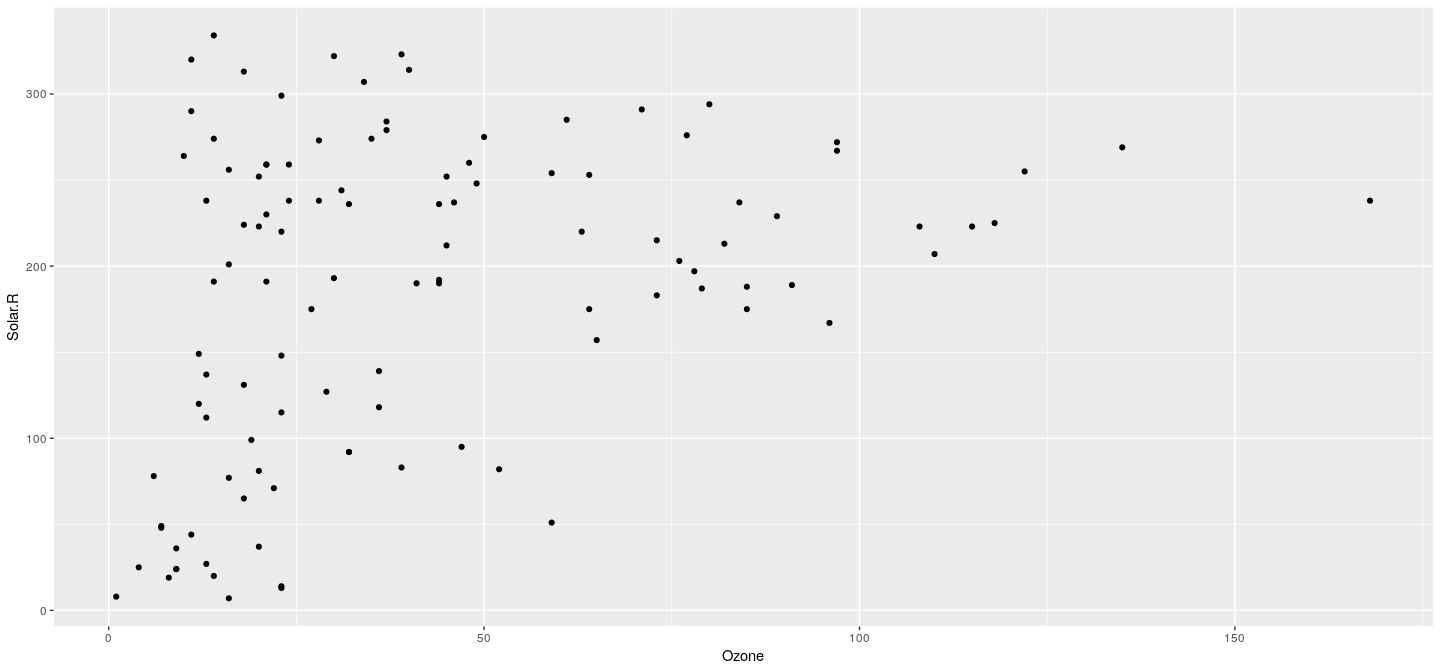
Example: scatter plot
- Equivalent call
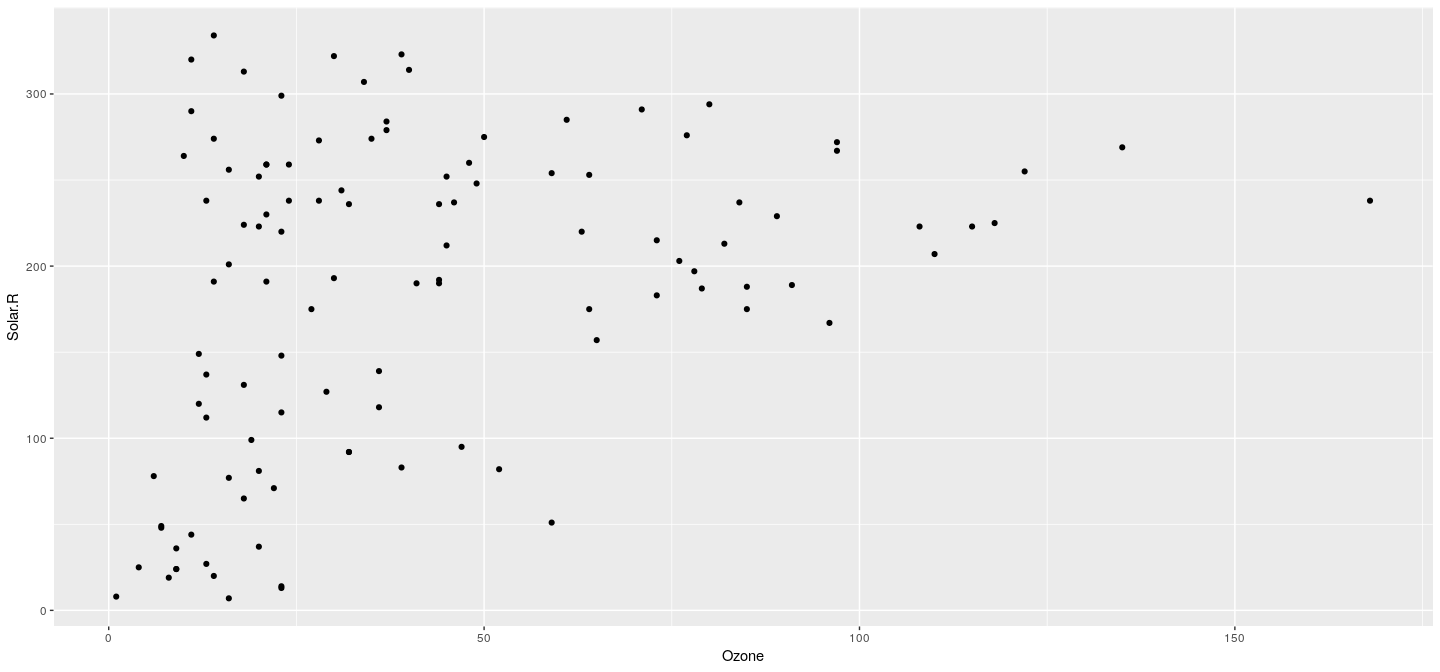
Example: scatter plot
- Something different (and meaningless)
p <- ggplot(airquality, aes(x = Ozone, y = Solar.R))
p + geom_bar(stat = "identity", position = "identity")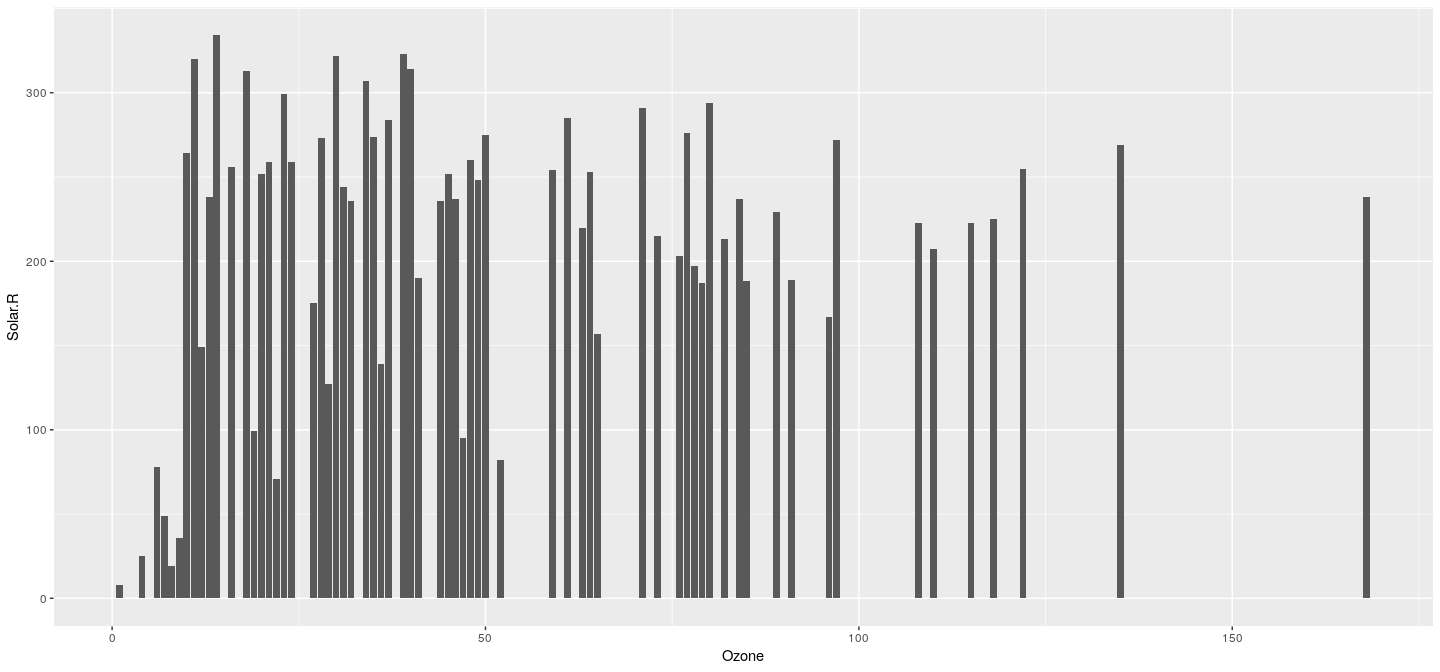
Example: scatter plot
The grammar approach makes it easy to create nonsense
But it also frees you from pre-defined plot types
Let’s go through some examples
Example: histogram
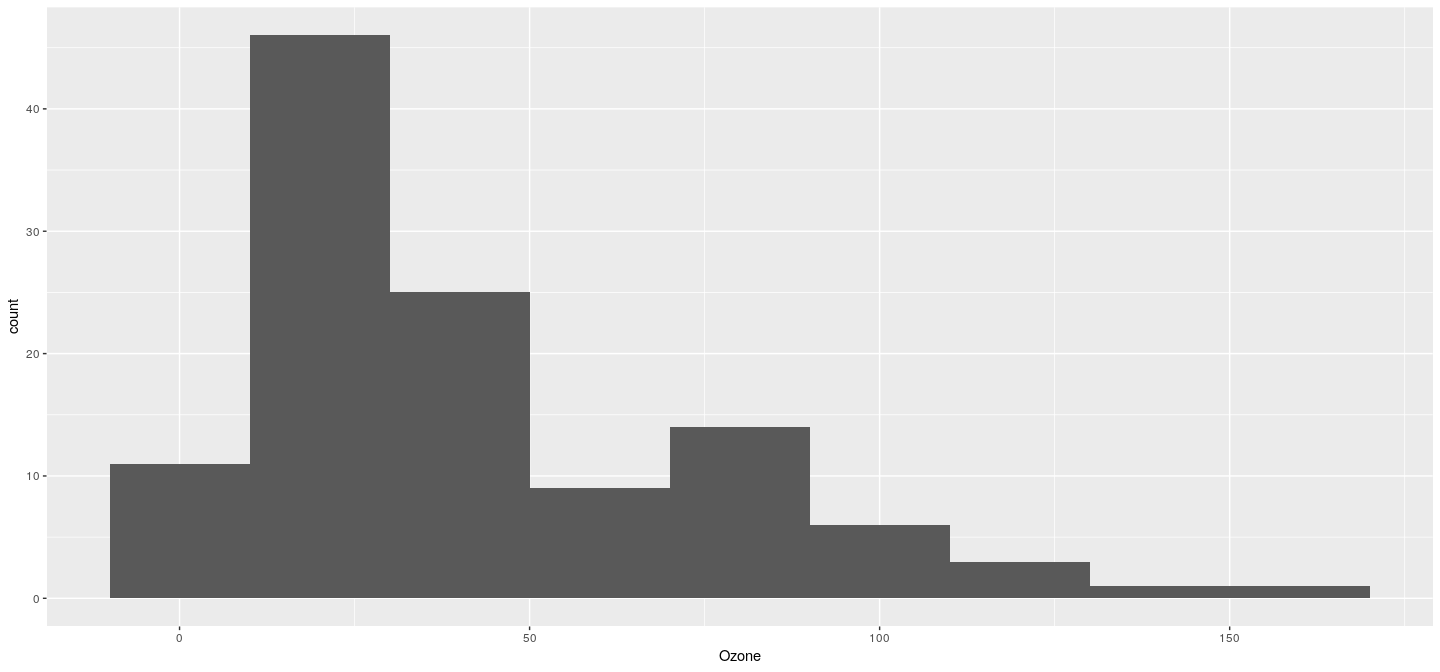
Example: density plot
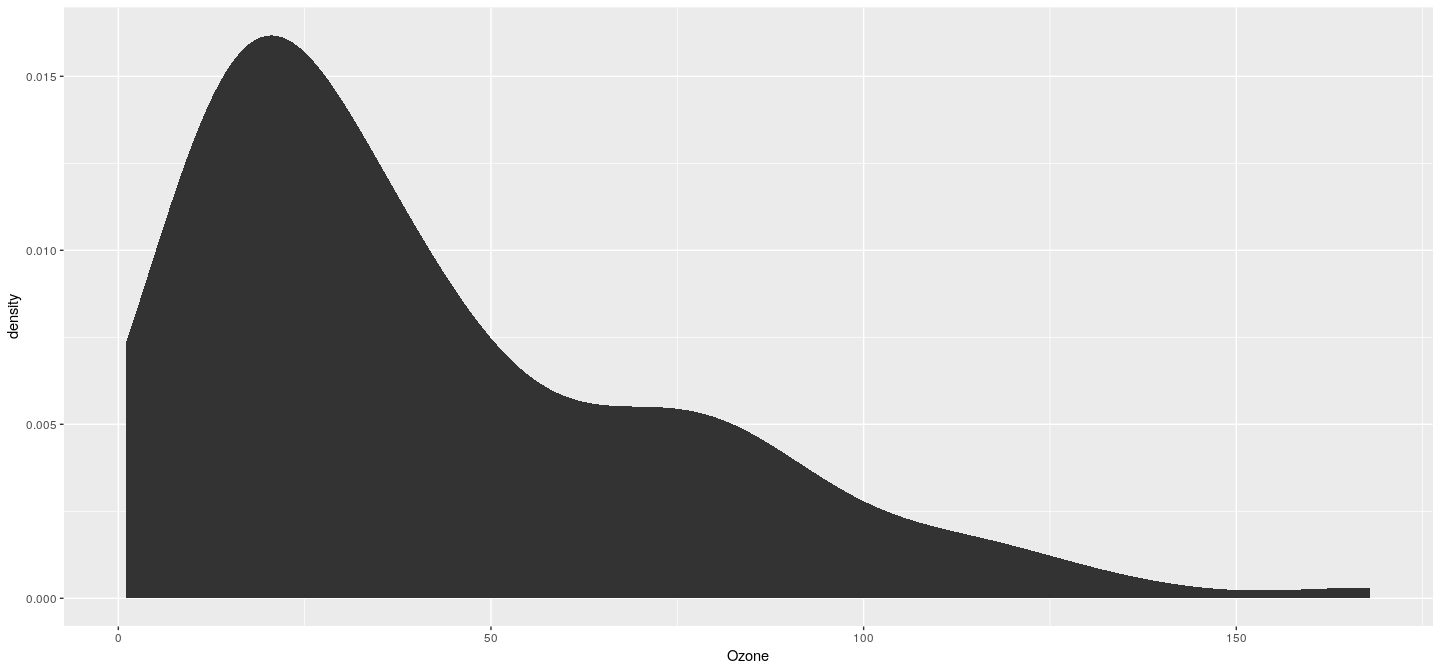
Example: density plot without shading
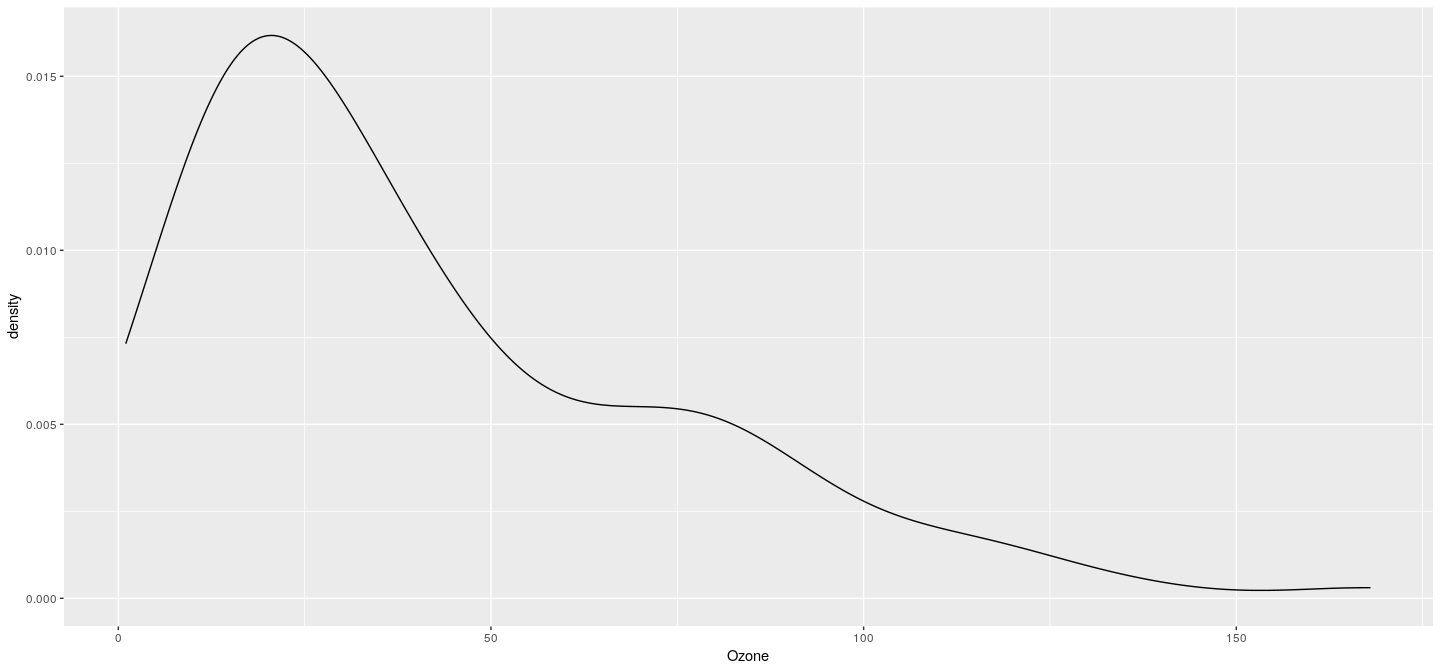
Example: density plot with groups
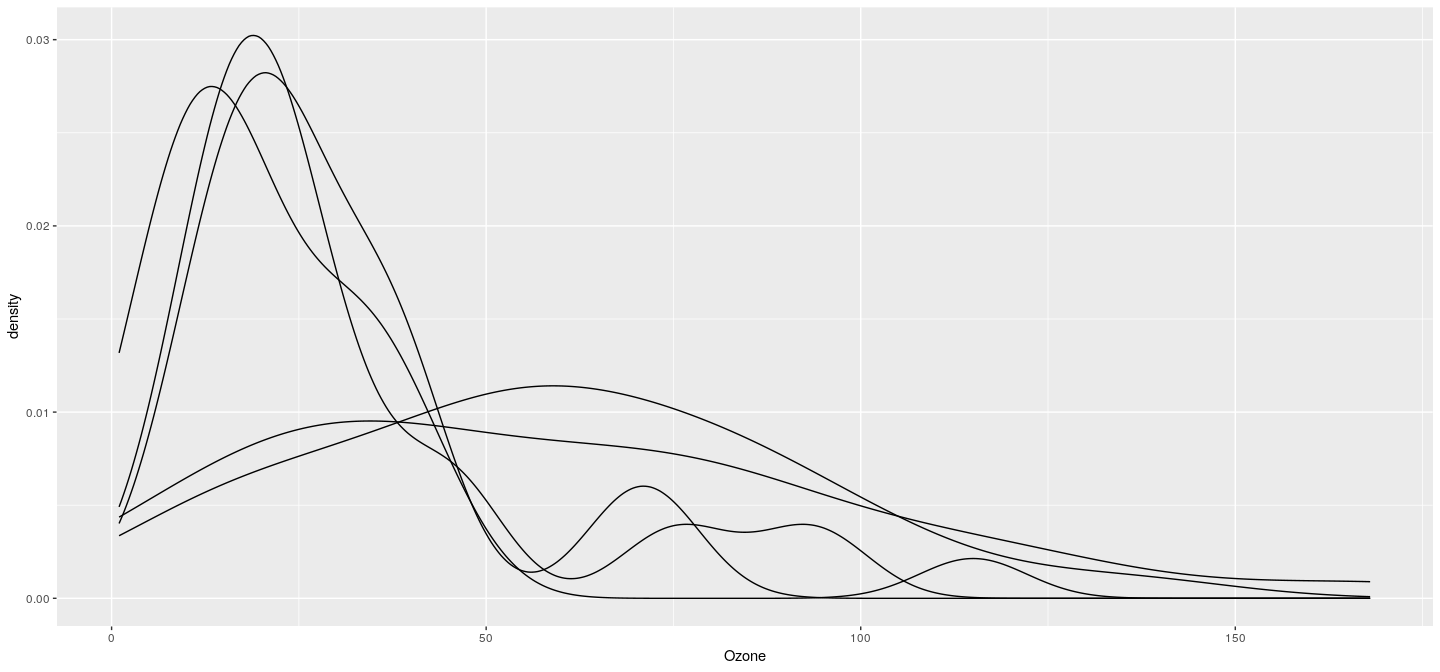
Example: density plot with colored groups
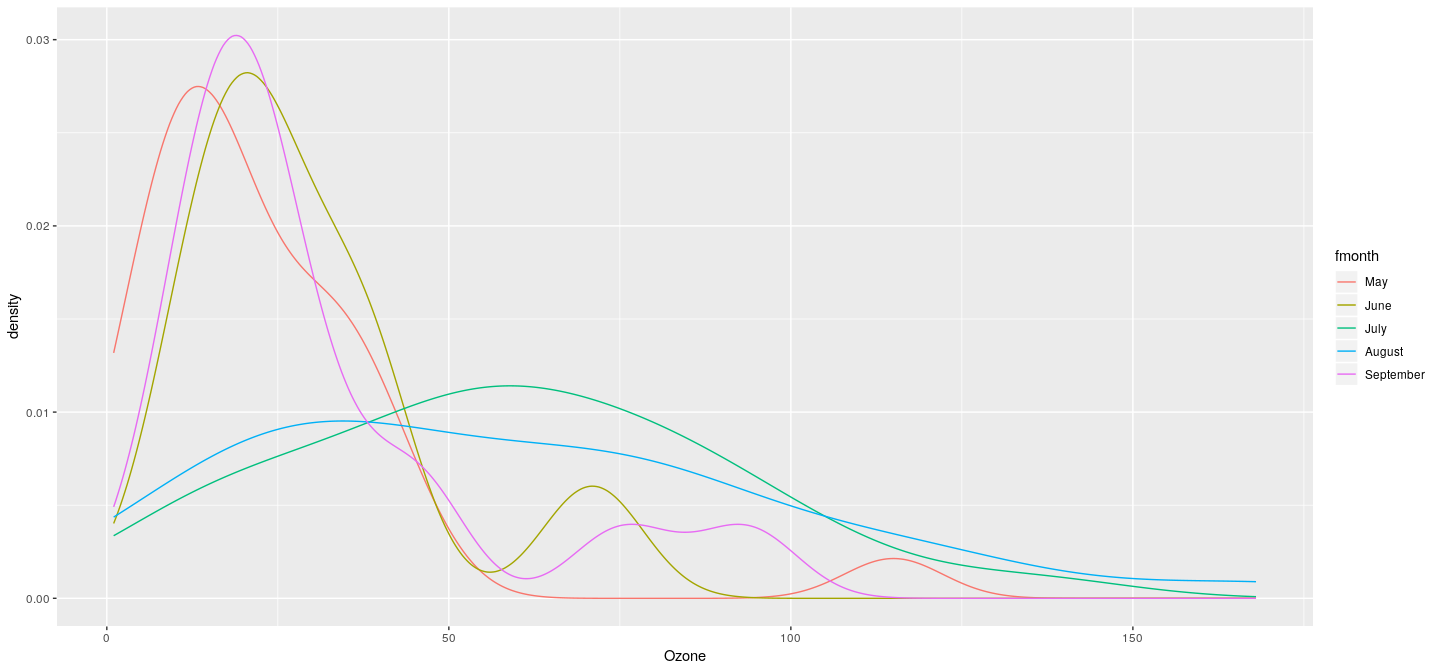
Example: density plot with faceting (conditioning)
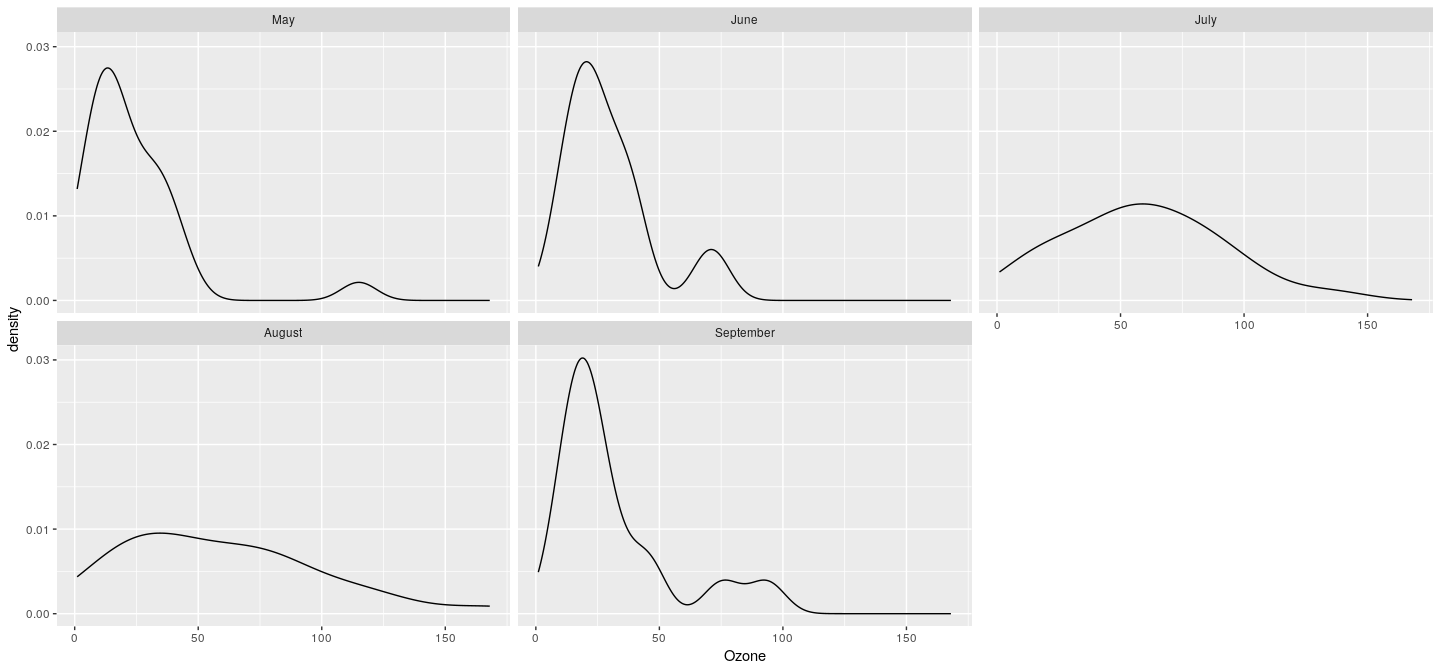
More compact calls using qplot()
Using the full grammar everytime is unnecessary
Most common plots can be created using
qplot()
Example: scatter plot
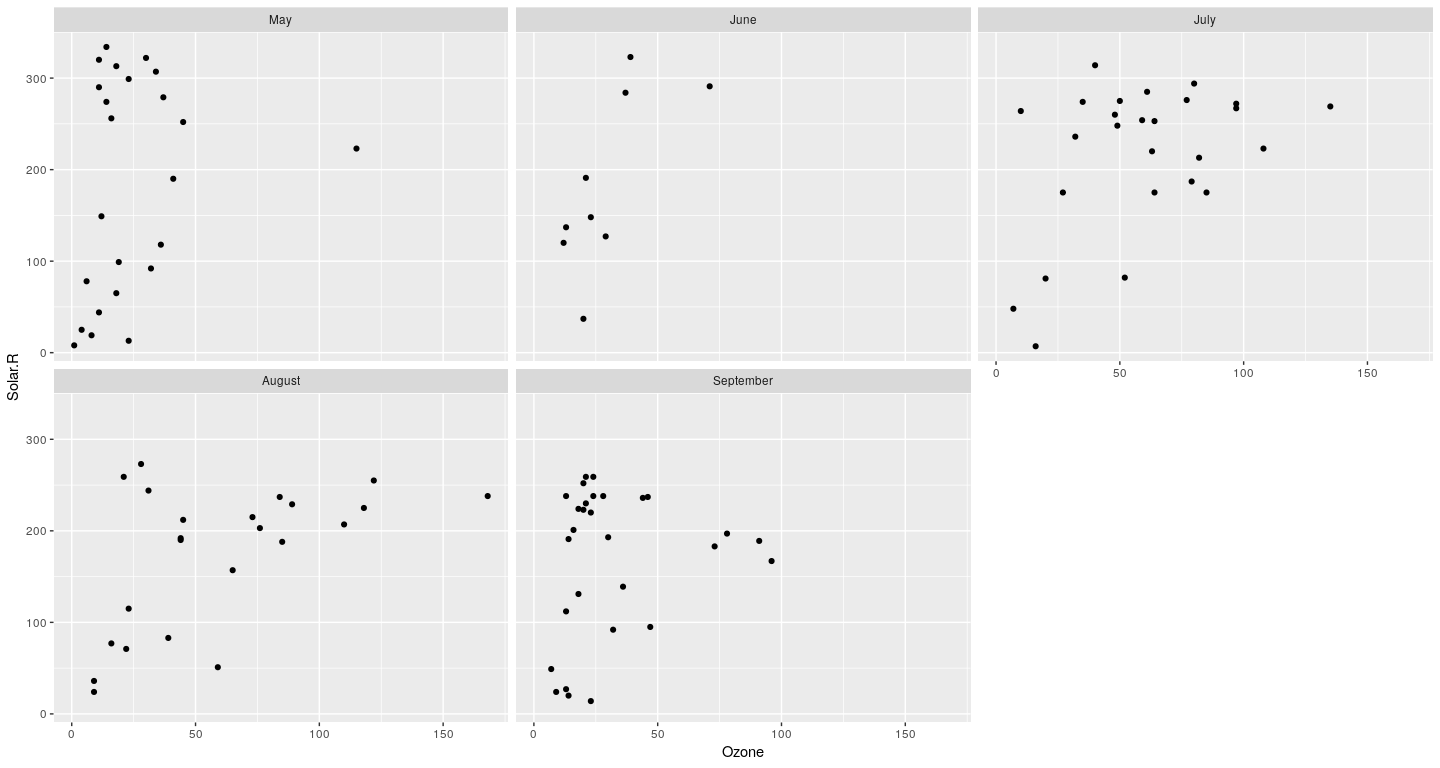
Example: density plot
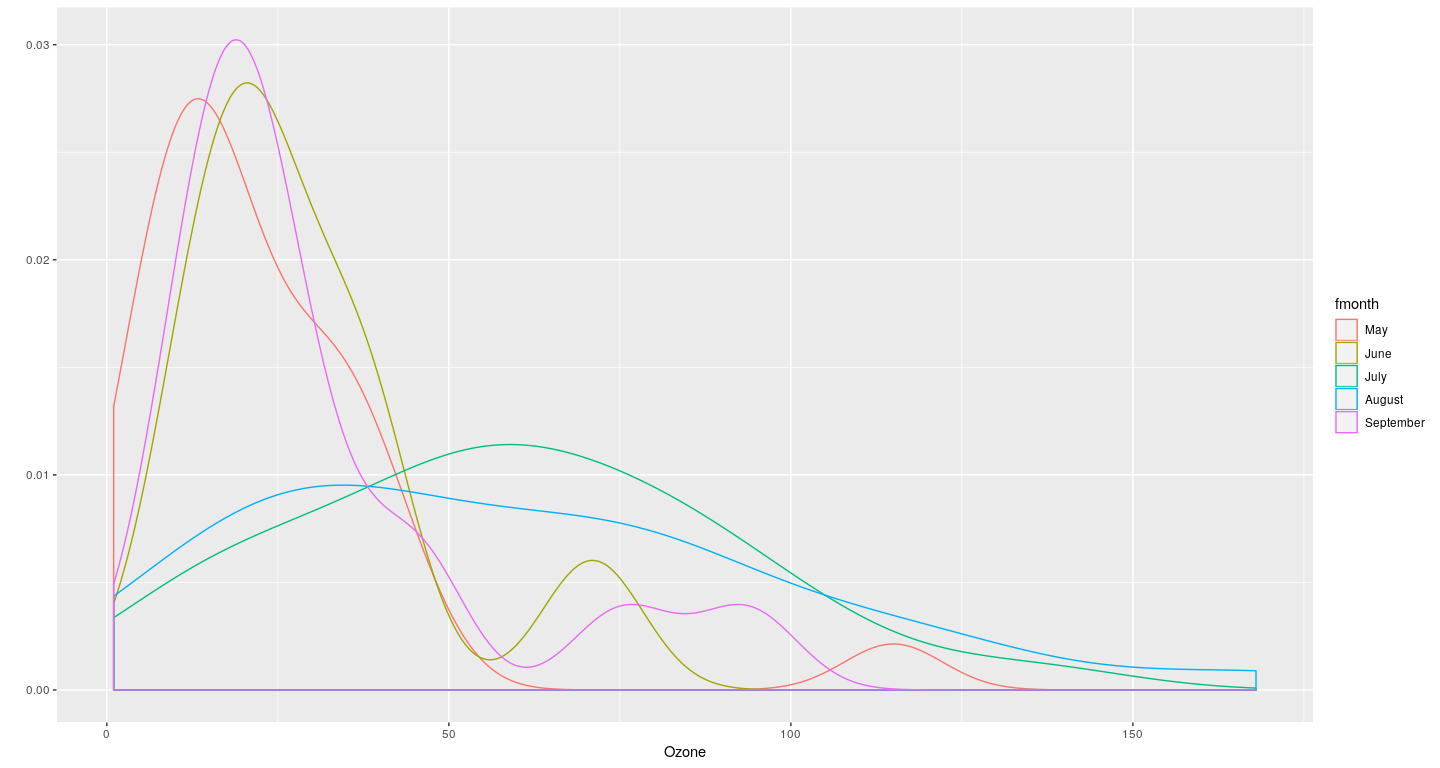
Layering to customize displays
qplot(x = x, y = y, data = anscombe.long, facets = ~ which) +
stat_smooth(method = "lm", se = FALSE)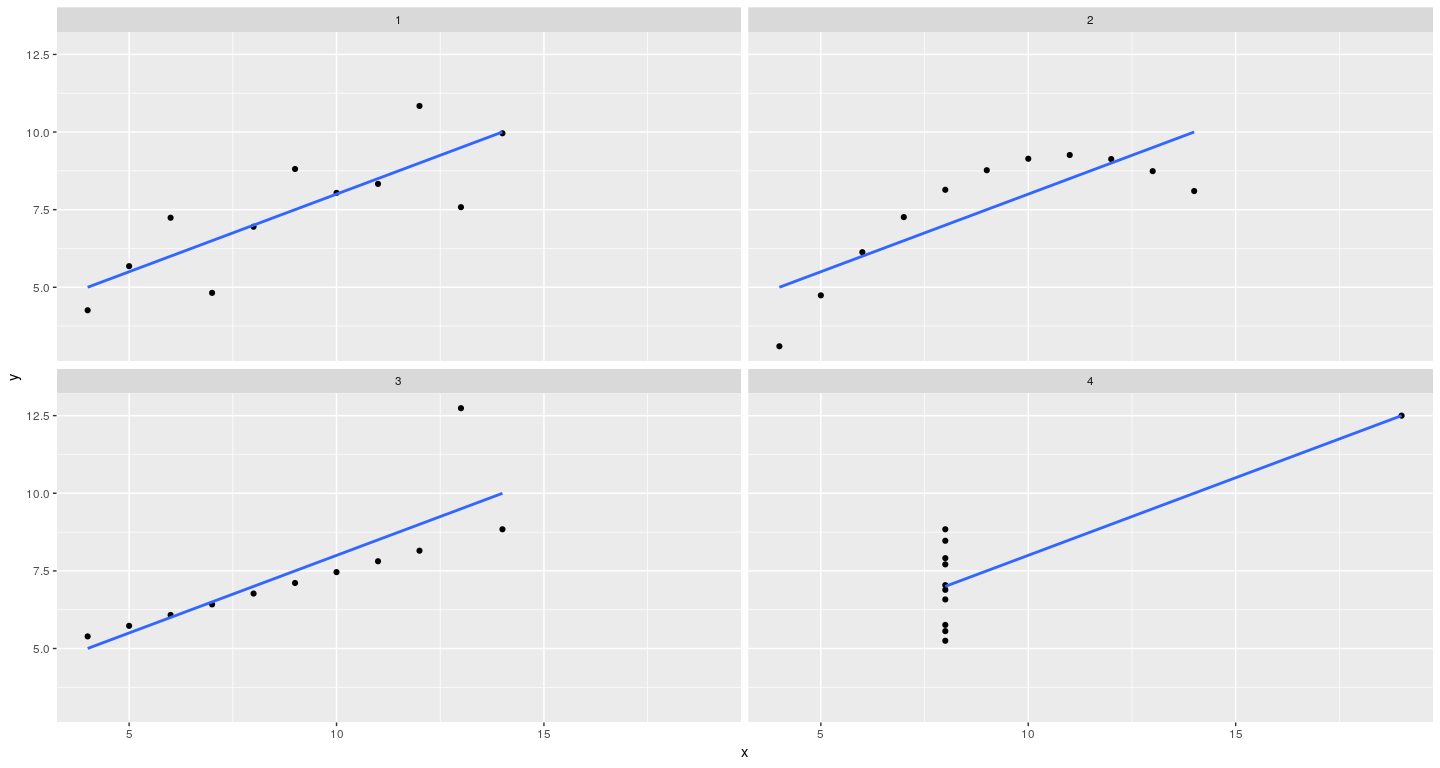
Layering to customize displays
qplot(x = x, y = y, data = anscombe.long, facets = ~ which) +
stat_smooth(method = "lm", se = FALSE) +
stat_smooth(method = "lm", formula = y ~ poly(x, 2), se = FALSE)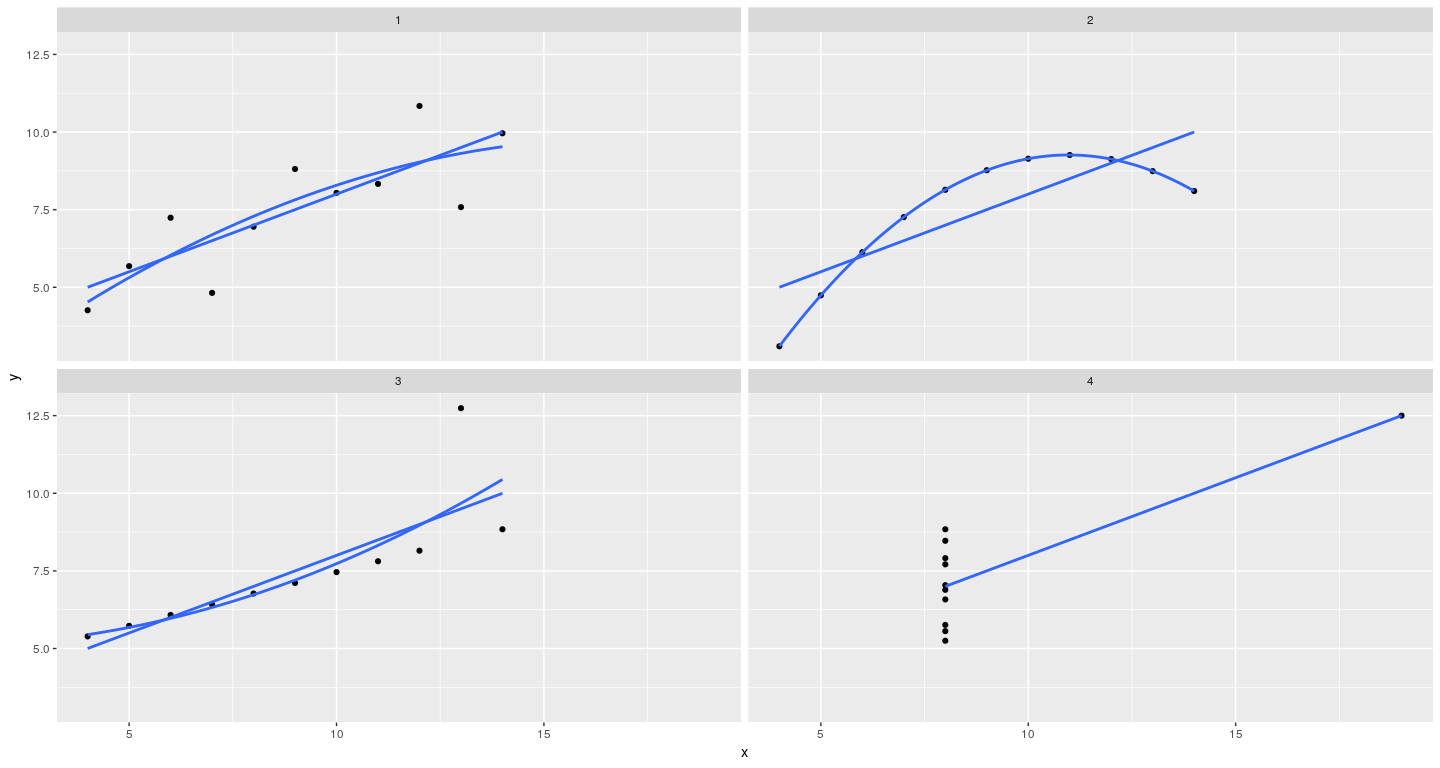
Dot plot
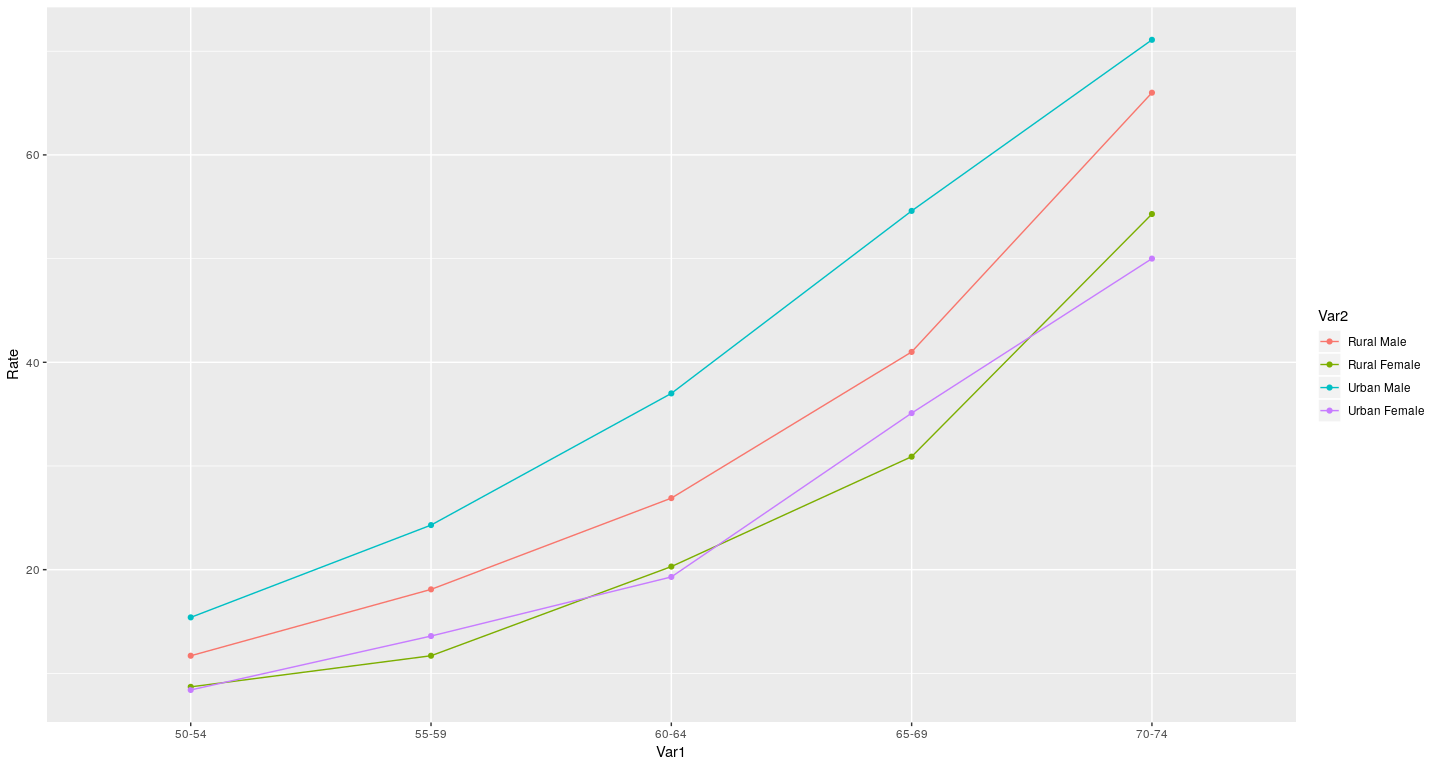
Interactive 3-D graphics
Traditional R graphics is usually good for static plots
There is some support for interaction, but this is rudimentary
A useful package for interactive 3-D plots is
rglUses
OpenGLto provide 3-D plots that can be rotated and zoomed
Interactive 3-D graphics using rgl
Further choices for interactive graphics
Several add-on packages provide some useful interfaces:
Package
plotlyandrbokeh: Javascript-based plots in browserPackage
rggobi: Interactive and dynamic graphics using GGobi (tours, brushing/linking)
Demos: