Static R Graphics: A Brief Introduction
Deepayan Sarkar
R graphics
R has a reputation for being a good system for graphics
This is mainly based on its ability to produce good publication-quality statistical plots
R actually has two largely independent graphics subsystems
Traditional graphics
- Available in R from the beginning
- Rich collection of tools
- Not very flexible
Grid graphics
- Relatively recent (2000)
- Low-level tool, highly flexible
Grid graphics, lattice and ggplot2
Grid graphics is not usually used directly by the user
But it forms the basis of two high-level graphics systems:
lattice: based on Trellis graphics (Cleveland)
ggplot2: inspired by “Grammar of Graphics” (Wilkinson)
These represent two very different philosophical approaches to graphics
lattice is in many ways a natural successor to traditional graphics
ggplot2 represents a completely different declarative approach
An example: Anscombe’s quartet
I will try to illustrate this with an example
Anscombe (1973) introduced four artificial bivariate datasets to emphasize the importance of graphics
The datasets all had the same means, standard deviations, and correlation
'data.frame': 11 obs. of 8 variables:
$ x1: num 10 8 13 9 11 14 6 4 12 7 ...
$ x2: num 10 8 13 9 11 14 6 4 12 7 ...
$ x3: num 10 8 13 9 11 14 6 4 12 7 ...
$ x4: num 8 8 8 8 8 8 8 19 8 8 ...
$ y1: num 8.04 6.95 7.58 8.81 8.33 ...
$ y2: num 9.14 8.14 8.74 8.77 9.26 8.1 6.13 3.1 9.13 7.26 ...
$ y3: num 7.46 6.77 12.74 7.11 7.81 ...
$ y4: num 6.58 5.76 7.71 8.84 8.47 7.04 5.25 12.5 5.56 7.91 ...An example: Anscombe’s quartet
x1 x2 x3 x4 y1 y2 y3 y4
9.000000 9.000000 9.000000 9.000000 7.500909 7.500909 7.500000 7.500909 x1 x2 x3 x4 y1 y2 y3 y4
3.316625 3.316625 3.316625 3.316625 2.031568 2.031657 2.030424 2.030579 [1] 0.8164205 0.8162365 0.8162867 0.8165214
- How can we plot all four datasets together?
Anscombe’s quartet using traditional graphics
Traditional graphics thinks of this as four different data sets
The function to create scatter plots is
plot()Multiple plots can be put in the same figure using
par(mfrow = ...)Several ways of specifying variable names inside dataset:
Anscombe’s quartet using traditional graphics
par(mfrow = c(1, 4))
plot(y1 ~ x1, data = anscombe, pch = 16)
plot(y2 ~ x2, data = anscombe, pch = 16)
plot(y3 ~ x3, data = anscombe, pch = 16)
plot(y4 ~ x4, data = anscombe, pch = 16)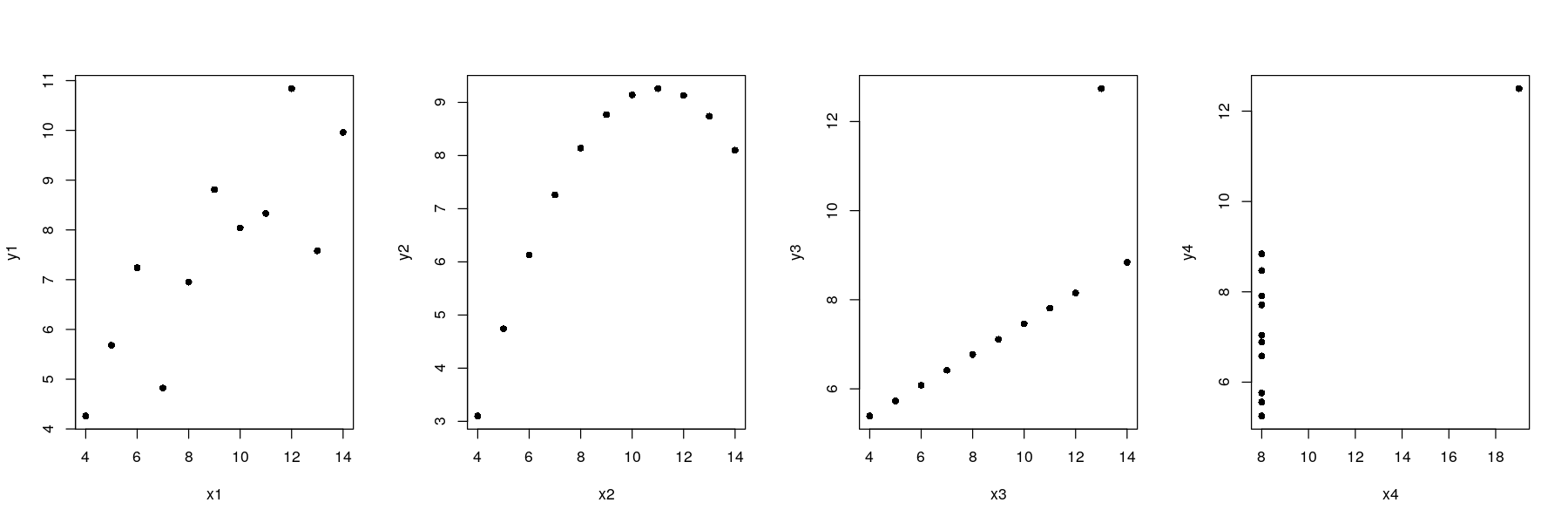
Anscombe’s quartet using traditional graphics
xrng <- with(anscombe, range(x1, x2, x3, x4))
yrng <- with(anscombe, range(y1, y2, y3, y4)) ## common axis limits
par(mfrow = c(1, 4))
plot(y1 ~ x1, data = anscombe, pch = 16, xlim = xrng, ylim = yrng)
plot(y2 ~ x2, data = anscombe, pch = 16, xlim = xrng, ylim = yrng)
plot(y3 ~ x3, data = anscombe, pch = 16, xlim = xrng, ylim = yrng)
plot(y4 ~ x4, data = anscombe, pch = 16, xlim = xrng, ylim = yrng)
Anscombe’s quartet using lattice and ggplot2
Both lattice and ggplot2 are capable of producing a single plot with all four datasets
But this requires the dataset to be in the “long format” (one row per data point)
anscombe.long <-
with(anscombe, data.frame(x = c(x1, x2, x3, x4),
y = c(y1, y2, y3, y4),
which = rep(c("1", "2", "3", "4"), each = 11)))
str(anscombe.long)'data.frame': 44 obs. of 3 variables:
$ x : num 10 8 13 9 11 14 6 4 12 7 ...
$ y : num 8.04 6.95 7.58 8.81 8.33 ...
$ which: Factor w/ 4 levels "1","2","3","4": 1 1 1 1 1 1 1 1 1 1 ...Anscombe’s quartet using lattice
library(package = "lattice")
xyplot(y ~ x | which, data = anscombe.long, layout = c(4, 1), pch = 16)
Anscombe’s quartet using ggplot2
library(package = "ggplot2")
ggplot(data = anscombe.long) + geom_point(aes(x = x, y = y)) + facet_grid( ~ which)
Anscombe’s quartet using lattice and ggplot2
The approaches share many common features
Both Capable of plotting subsets of data (indexed by categorical variables)
- This idea is known by several names: small multiples, conditioning, facetting
Both makes efficient use of available space (common scales, common axes)
Different visual appearance, but that is superficial (different default themes)
Anscombe’s quartet using lattice and ggplot2
However, the way in which we specify the display is very different
lattice uses an extension of the formula-data interface (with function
xyplot()instead ofplot())ggplot2 specifies type of rendering (geom) and mapping of variables to coordinates (aesthetics)
The differences become clearer if we try to customize the display further
A natural modification in this example is to add a linear regression line to each scatter plot
Anscombe’s quartet with regression lines
xrng <- with(anscombe, range(x1, x2, x3, x4))
yrng <- with(anscombe, range(y1, y2, y3, y4))
par(mfrow = c(1, 4))
plot(y1 ~ x1, anscombe, pch = 16, xlim = xrng, ylim = yrng)
abline(lm(y1 ~ x1, anscombe)) ## lm() fits regression line, abline() adds the line to current plot
plot(y2 ~ x2, anscombe, pch = 16, xlim = xrng, ylim = yrng)
abline(lm(y2 ~ x2, anscombe))
plot(y3 ~ x3, anscombe, pch = 16, xlim = xrng, ylim = yrng)
abline(lm(y3 ~ x3, anscombe))
plot(y4 ~ x4, anscombe, pch = 16, xlim = xrng, ylim = yrng)
abline(lm(y4 ~ x4, anscombe))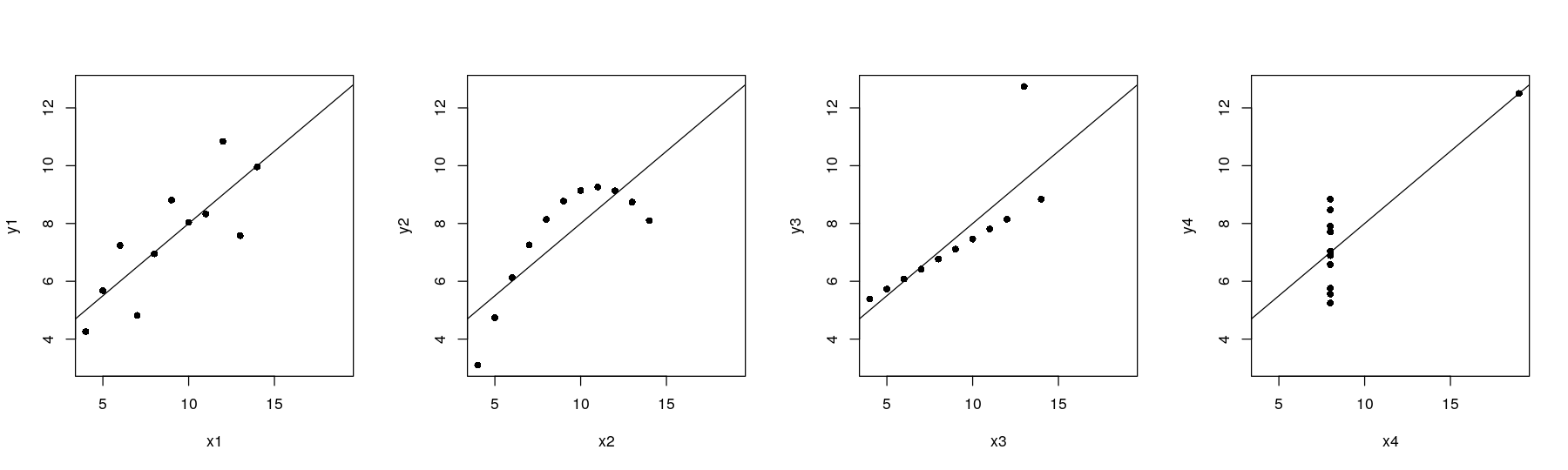
Anscombe’s quartet with regression lines
The traditional graphics approach is to add the line after the plot is drawn
In general, a plot is never finished, you can always add more points, lines, text, …
This is possible because there is only one plot !
lattice and ggplot2 need alternative solutions
The ggplot2 solution is to allow plots to have multiple layers
The lattice solution is to allow user to fully specify the procedure used to display data
Regression lines: the ggplot2 solution
- Plot with points only
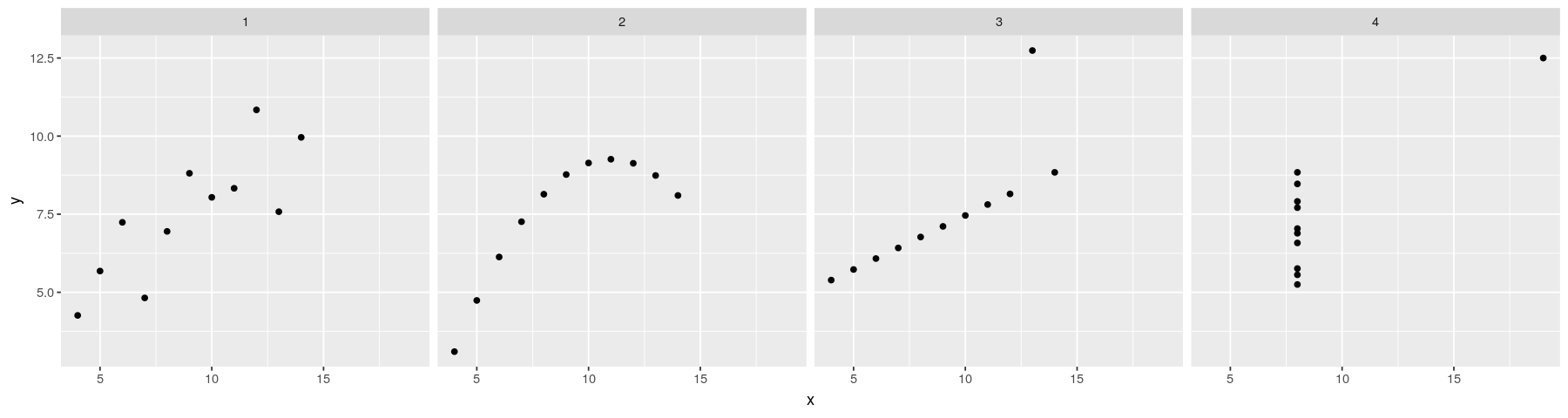
Regression lines: the ggplot2 solution
- Plot with regression line only
ggplot(data = anscombe.long) + facet_grid( ~ which) +
geom_smooth(aes(x = x, y = y), method = "lm", se = FALSE)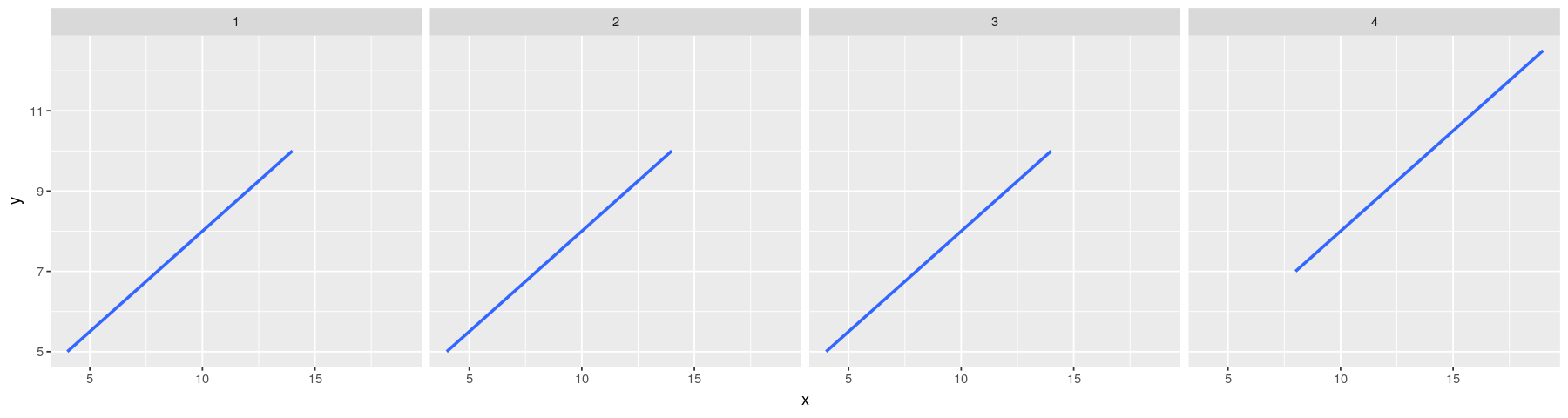
Regression lines: the ggplot2 solution
- Plot with both points and regression lines
ggplot(data = anscombe.long) + facet_grid( ~ which) + geom_point(aes(x = x, y = y)) +
geom_smooth(aes(x = x, y = y), method = "lm", se = FALSE)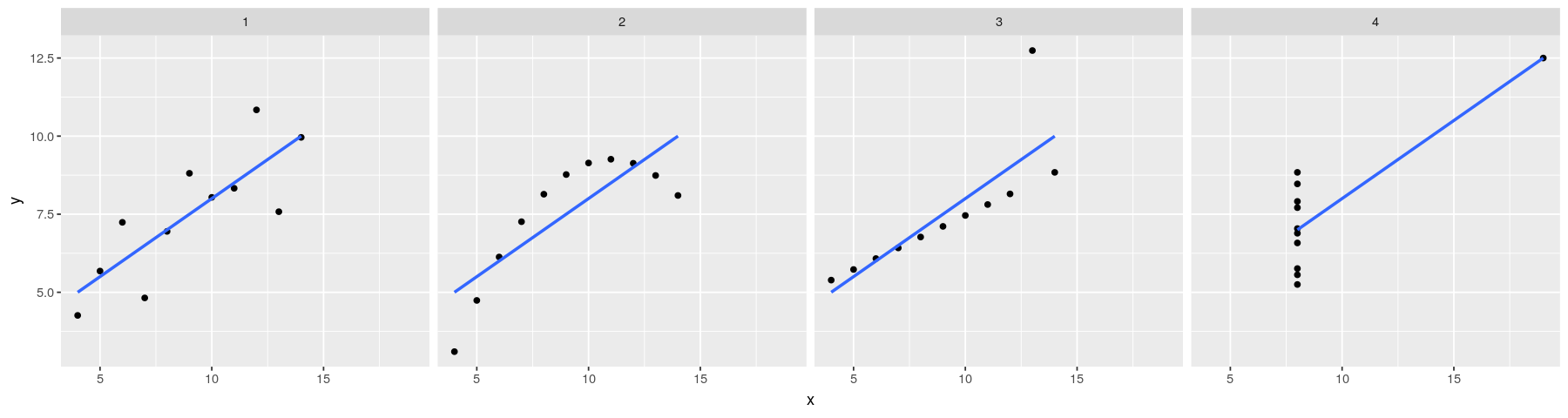
Regression lines: the lattice solution
The lattice solution is actually very similar to the traditional graphics solution
Basically, we want to do the following for each data subset
x, y:Draw points at
(x, y)Draw the linear regression line through
x, y
For lattice, we need to encapsulate this procedure into a function
displayFunction <- function(x, y) {
panel.grid(h = -1, v = -1) ## add a reference grid
panel.points(x, y, pch = 16) ## draw the points
panel.abline(lm(y ~ x), col = "grey50") ## draw linear regression line
}- This function is then supplied to the
xyplot()function as thepanelargument
Regression lines: the lattice solution
- Plot with points only
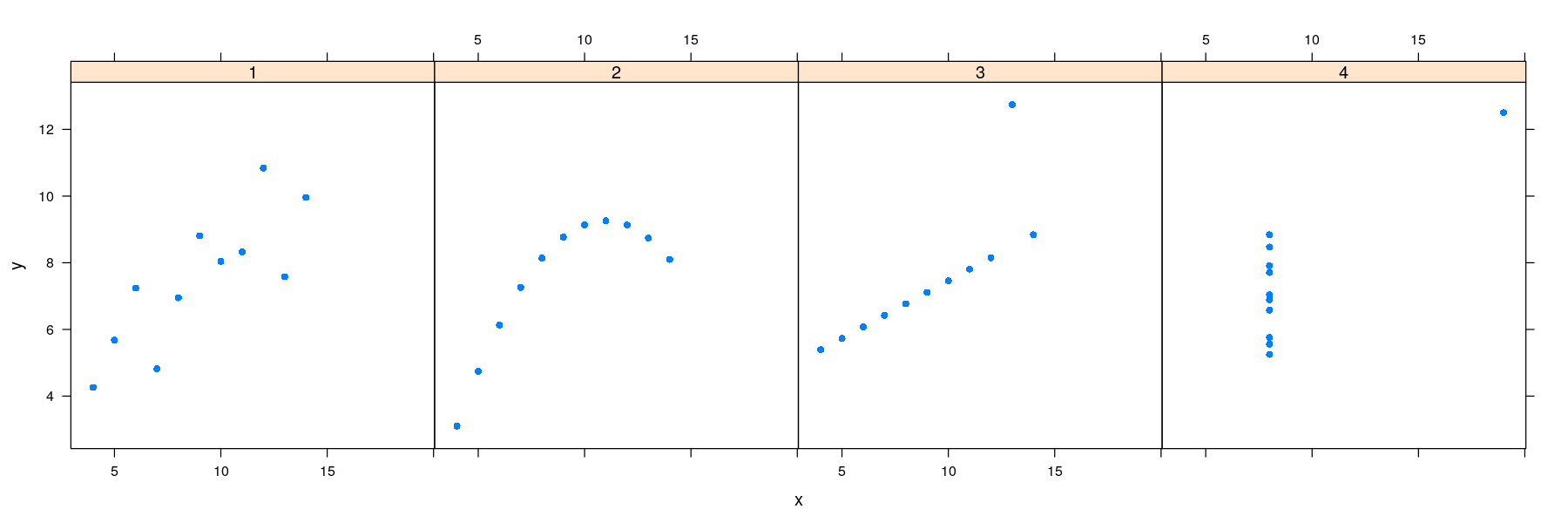
Regression lines: the lattice solution
- Plot with grid, points, and regression line

Regression lines: the lattice solution
- lattice also supports a layering mechanism similar to ggplot2
library(package = "latticeExtra")
xyplot(y ~ x | which, data = anscombe.long, layout = c(4, 1), pch = 16) +
layer_(panel.grid(h = -1, v = -1)) + layer(panel.abline(lm(y ~ x)))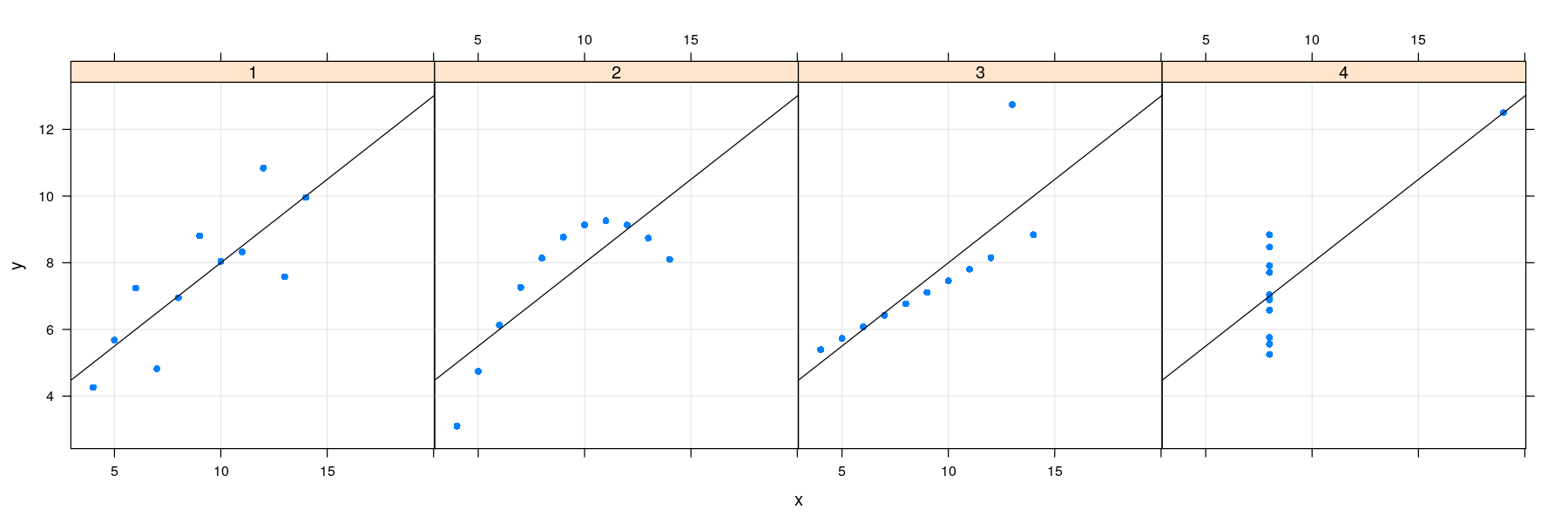
Regression lines: the lattice solution
- Common customizations like these are also supported directly through optional arguments
library(package = "latticeExtra")
xyplot(y ~ x | which, data = anscombe.long, layout = c(4, 1), pch = 16,
grid = TRUE, type = c("p", "r"))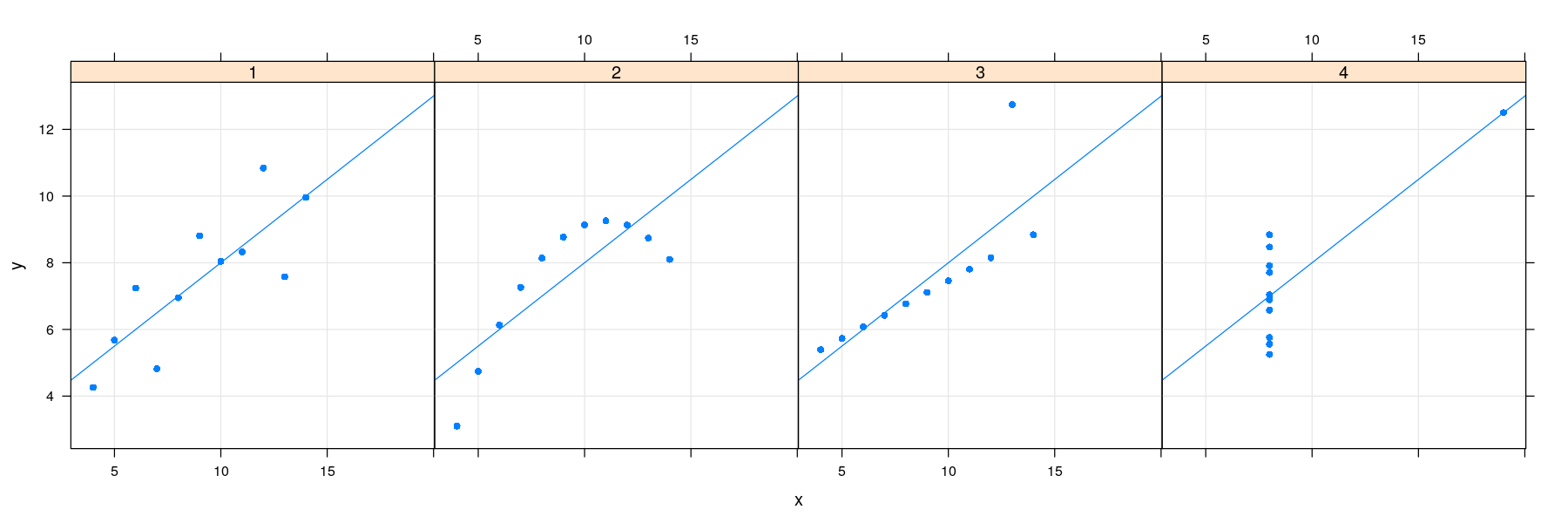
Brief history of R graphics
R is an Open Source / Free Software re-implementation / dialect of S
Freely available from CRAN
Available on all major platforms (Windows / UNIX / Linux / Mac)
The original implementation, available commercially as S-PLUS, was developed at Bell Labs
Both traditional graphics and Trellis graphics were part of the original S
Remembering this helps in understanding the design of graphics in R
Origins of the graphics model
The abstract graphics model in S may be described as a “painter’s model”:
a graphic is built out of a small set of primitives such as line segments, polygons, text, etc., and
later elements are drawn on top of earlier ones
no provision for deleting an element once it was drawn
except to start a completely new graphic
Advantage: both input and output could be abstracted
Graphics functions called by users would internally call these primitives
For output, the primitives could be implemented differently depending on the target “device”
This leads to a concept of graphics devices
Postscript or PDF files for printing,
Hardware devices such as pen plotters,
On-screen devices for interactive viewing (different for Windows / Linux / Mac)
Image files (JPG / PNG) for inclusion in a web page
Device-specific implementations of the primitives are known as device drivers
New drivers can be written to support new kinds of output formats
See
?Devicesfor more details.This is how R has solved the problem of cross-platform consistency for graphics
The drawback is that more advanced / interactive features are not available
Traditional graphics
The core of the traditional R graphics system is the suite of functions available in the graphics package,
Various add-on packages providing further functionality.
The full list of functions can be seen using
The listed functions can be roughly categorized into two groups:
High-level functions are intended to produce a complete plot by themselves
Low-level functions are intended to add elements to existing plots
Commonly used high-level traditional graphics functions
| Function | Default Display |
|---|---|
plot() |
Scatter Plot, Time-series Plot (with type="l") |
boxplot() |
Comparative Box-and-Whisker Plots |
barplot() |
Bar Plot |
dotchart() |
Cleveland Dot Plot |
hist() |
Histogram |
plot(density()) |
Kernel Density Plot |
qqnorm() |
Normal Quantile-Quantile Plot |
qqplot() |
Two-sample Quantile-Quantile Plot |
stripchart() |
Stripchart (Comparative 1-D Scatter Plots) |
pairs() |
Scatter-Plot Matrix |
lattice defines analogous functions with different names
| Function | Default Display |
|---|---|
xyplot() |
Scatter Plot, Time-series Plot (with type="l") |
bwplot() |
Comparative Box-and-Whisker Plots |
barchart() |
Bar Plot |
dotplot() |
Cleveland Dot Plot |
histogram() |
Histogram |
densityplot() |
Kernel Density Plot |
qqmath() |
Normal Quantile-Quantile Plot |
qq() |
Two-sample Quantile-Quantile Plot |
stripplot() |
Stripchart (Comparative 1-D Scatter Plots) |
splom() |
Scatter-Plot Matrix |
Grammar of graphics
Traditional and trellis graphics both have the same basic approach:
Functions are written to implement specific graphical designs
Usually these are designs that have been established as being useful
Customization is achieved through a procedural approach
The ggplot2 package takes a different declarative approach:
Defines a “layered grammar” for defining graphical designs
Final display is a composition of various components
A systematic grammar is used to specify the composition
Can be used to create novel displays easily
Plots consist of one or more layers (e.g., raw data could be one layer, model fits another)
Grammar of graphics: Main components
Aesthetic mappings that map data values to some aspect in the displayed graph, such as
- coordinate positions
- color, shape, size
- group, …
geometric types used to render the mapped data, e.g.,
- points, lines, polygons
- more complex types such as a box-and-whisker plot
statistical transformations that are applied to the data beforehand, such as
- binning for histograms
- computation of kernel density estimates.
Scales that give a visual indication of the aesthetic mappings, e.g.,
- axis annotation for position mapping
- legends for mapping to color, size, etc.
Faceting (conditioning) to produce small multiples
Grammar of graphics - built-in geoms and stats
- Geoms
[1] "geom_abline" "geom_area" "geom_bar" "geom_bin2d" "geom_blank" "geom_boxplot"
[7] "geom_col" "geom_contour" "geom_count" "geom_crossbar" "geom_curve" "geom_density"
[13] "geom_density_2d" "geom_density2d" "geom_dotplot" "geom_errorbar" "geom_errorbarh" "geom_freqpoly"
[19] "geom_hex" "geom_histogram" "geom_hline" "geom_jitter" "geom_label" "geom_line"
[25] "geom_linerange" "geom_map" "geom_path" "geom_point" "geom_pointrange" "geom_polygon"
[31] "geom_qq" "geom_qq_line" "geom_quantile" "geom_raster" "geom_rect" "geom_ribbon"
[37] "geom_rug" "geom_segment" "geom_sf" "geom_sf_label" "geom_sf_text" "geom_smooth"
[43] "geom_spoke" "geom_step" "geom_text" "geom_tile" "geom_violin" "geom_vline"
- Stats
[1] "stat_bin" "stat_bin_2d" "stat_bin_hex" "stat_bin2d" "stat_binhex"
[6] "stat_boxplot" "stat_contour" "stat_count" "stat_density" "stat_density_2d"
[11] "stat_density2d" "stat_ecdf" "stat_ellipse" "stat_function" "stat_identity"
[16] "stat_qq" "stat_qq_line" "stat_quantile" "stat_sf" "stat_sf_coordinates"
[21] "stat_smooth" "stat_spoke" "stat_sum" "stat_summary" "stat_summary_2d"
[26] "stat_summary_bin" "stat_summary_hex" "stat_summary2d" "stat_unique" "stat_ydensity" Parting comments
We don’t have time to go into details of any of these systems
Lots of help easily available on the internet
If you primarily use R / Python for your analysis, I would also suggest learning about
Together, these form a convenient basis for “literate documents” combining text and code
This talk is an example
Very good support available in R Studio, the best interface to R for beginners
References
Anscombe, Francis J. 1973. “Graphs in Statistical Analysis.” The American Statistician 27 (1). Taylor & Francis Group: 17–21.